The effect of SNCA 3' region on the levels of SNCA-112 splicing variant
- PMID: 21046180
- PMCID: PMC3030669
- DOI: 10.1007/s10048-010-0263-4
The effect of SNCA 3' region on the levels of SNCA-112 splicing variant
Abstract
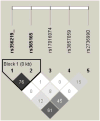
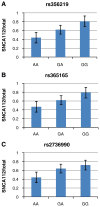
Genetic variability at the 3' region of SNCA locus has been repeatedly associated with susceptibility to sporadic Parkinson's disease (PD). Accumulated evidence emphasizes the importance of SNCA dosage and expression levels in PD pathogenesis. However, the mechanism through which the 3' region of SNCA gene modulates the risk to develop sporadic PD remained elusive. We studied the effect of PD risk-associated variants at SNCA 3' regions on SNCA112-mRNA (exon 5 in-frame skipping) levels in vivo in 117 neuropathologically normal, human brain frontal cortex samples. SNPs tagging the SNCA 3' showed significant effects on the relative levels of SNCA112-mRNA from total SNCA transcripts levels. The "risk" alleles were correlated with increased expression ratio of SNCA112-mRNA from total. We provide evidence for functional consequences of PD-associated SNCA gene variants at the 3' region, suggesting that genetic regulation of SNCA splicing plays an important role in the development of the disease. Further studies to determine the definite functional variant/s within SNCA 3'and to establish their association with PD pathology are necessary.
Conflict of interest statement
Figures
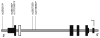
Similar articles
-
Genetic regulation of alpha-synuclein mRNA expression in various human brain tissues.PLoS One. 2009 Oct 16;4(10):e7480. doi: 10.1371/journal.pone.0007480. PLoS One. 2009. PMID: 19834617 Free PMC article.
-
SNCA variants and alpha-synuclein level in CD45+ blood cells in Parkinson's disease.J Neurol Sci. 2018 Dec 15;395:135-140. doi: 10.1016/j.jns.2018.10.002. Epub 2018 Oct 3. J Neurol Sci. 2018. PMID: 30316070
-
SNCA 3' UTR Genetic Variants in Patients with Parkinson's Disease.Biomolecules. 2021 Nov 30;11(12):1799. doi: 10.3390/biom11121799. Biomolecules. 2021. PMID: 34944443 Free PMC article.
-
Genetic variability in SNCA and Parkinson's disease.Neurogenetics. 2011 Nov;12(4):283-93. doi: 10.1007/s10048-011-0292-7. Epub 2011 Jul 29. Neurogenetics. 2011. PMID: 21800132 Review.
-
Association between alpha-synuclein (SNCA) rs11931074 variability and susceptibility to Parkinson's disease: an updated meta-analysis of 41,811 patients.Neurol Sci. 2020 Feb;41(2):271-280. doi: 10.1007/s10072-019-04107-8. Epub 2019 Nov 22. Neurol Sci. 2020. PMID: 31758346 Review.
Cited by
-
Postmortem Interval Influences α-Synuclein Expression in Parkinson Disease Brain.Parkinsons Dis. 2012;2012:614212. doi: 10.1155/2012/614212. Epub 2012 Mar 13. Parkinsons Dis. 2012. PMID: 22530163 Free PMC article.
-
Head injury, α-synuclein genetic variability and Parkinson's disease.Eur J Neurol. 2015 May;22(5):874-8. doi: 10.1111/ene.12585. Epub 2014 Nov 5. Eur J Neurol. 2015. PMID: 25370538 Free PMC article.
-
iPS Cell-Based Model for MAPT Haplotype as a Risk Factor for Human Tauopathies Identifies No Major Differences in TAU Expression.Front Cell Dev Biol. 2021 Aug 31;9:726866. doi: 10.3389/fcell.2021.726866. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34532319 Free PMC article.
-
Orchestrated increase of dopamine and PARK mRNAs but not miR-133b in dopamine neurons in Parkinson's disease.Neurobiol Aging. 2014 Oct;35(10):2302-15. doi: 10.1016/j.neurobiolaging.2014.03.016. Epub 2014 Mar 22. Neurobiol Aging. 2014. PMID: 24742361 Free PMC article.
-
Analysis of Genetic and Non-genetic Predictors of Levodopa Induced Dyskinesia in Parkinson's Disease.Front Pharmacol. 2021 Apr 29;12:640603. doi: 10.3389/fphar.2021.640603. eCollection 2021. Front Pharmacol. 2021. PMID: 33995045 Free PMC article.
References
-
- Polymeropoulos MH, Lavedan C, Leroy E, Ide SE, Dehejia A, et al. Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease. Science. 1997;276:2045–2047. - PubMed
-
- Singleton AB, Farrer M, Johnson J, Singleton A, Hague S, et al. alpha-Synuclein locus triplication causes Parkinson’s disease. Science. 2003;302:841. - PubMed
-
- Farrer M, Kachergus J, Forno L, Lincoln S, Wang DS, et al. Comparison of kindreds with parkinsonism and alpha-synuclein genomic multiplications. Ann Neurol. 2004;55:174–179. - PubMed
-
- Miller DW, Hague SM, Clarimon J, Baptista M, Gwinn-Hardy K, et al. Alpha-synuclein in blood and brain from familial Parkinson disease with SNCA locus triplication. Neurology. 2004;62:1835–1838. - PubMed
-
- Chartier-Harlin MC, Kachergus J, Roumier C, Mouroux V, Douay X, et al. Alpha-synuclein locus duplication as a cause of familial Parkinson’s disease. Lancet. 2004;364:1167–1169. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

