Replication factor C complexes play unique pro- and anti-establishment roles in sister chromatid cohesion
- PMID: 21060875
- PMCID: PMC2965161
- DOI: 10.1371/journal.pone.0015381
Replication factor C complexes play unique pro- and anti-establishment roles in sister chromatid cohesion
Abstract
Recent studies have lead to a rapid expansion of sister chromatid cohesion pathways. Of particular interest is the growth in classifications of anti-establishment factors-now including those that are cohesin-associated (Rad61/WAPL and Pds5) or DNA replication fork-associated (Elg1-RFC). In this study, we show that the two classes of anti-establishment complexes are indistinguishable when challenged both genetically and functionally. These findings suggest that both classes function in a singular pathway that is centered on Ctf7/Eco1 (herein termed Ctf7) regulation. The anti-establishment activity of Elg1-RFC complex is particular intriguing given that an alternate Ctf18-RFC complex exhibits robust pro-establishment activity. Here, we provide several lines of evidence, including the use of Ctf7 bypass suppressors, indicating that these activities are not simply antagonistic. Moreover, the results suggest that Ctf18-RFC is capable of promoting sister chromatid pairing reactions independent of Ctf7. The combination of these studies suggest a new model of sister chromatid pairing regulation.
Conflict of interest statement
Figures


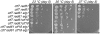



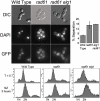
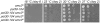

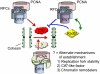
Similar articles
-
The Elg1-RFC clamp-loading complex performs a role in sister chromatid cohesion.PLoS One. 2009;4(3):e4707. doi: 10.1371/journal.pone.0004707. Epub 2009 Mar 5. PLoS One. 2009. PMID: 19262753 Free PMC article.
-
Division of Labor between PCNA Loaders in DNA Replication and Sister Chromatid Cohesion Establishment.Mol Cell. 2020 May 21;78(4):725-738.e4. doi: 10.1016/j.molcel.2020.03.017. Epub 2020 Apr 10. Mol Cell. 2020. PMID: 32277910 Free PMC article.
-
Rfc5p regulates alternate RFC complex functions in sister chromatid pairing reactions in budding yeast.Cell Cycle. 2010 Nov 1;9(21):4370-8. doi: 10.4161/cc.9.21.13634. Epub 2010 Nov 13. Cell Cycle. 2010. PMID: 20980821 Free PMC article.
-
Integrating Sister Chromatid Cohesion Establishment to DNA Replication.Genes (Basel). 2022 Mar 31;13(4):625. doi: 10.3390/genes13040625. Genes (Basel). 2022. PMID: 35456431 Free PMC article. Review.
-
The Interplay of Cohesin and the Replisome at Processive and Stressed DNA Replication Forks.Cells. 2021 Dec 8;10(12):3455. doi: 10.3390/cells10123455. Cells. 2021. PMID: 34943967 Free PMC article. Review.
Cited by
-
G1-Cyclin2 (Cln2) promotes chromosome hypercondensation in eco1/ctf7 rad61 null cells during hyperthermic stress in Saccharomyces cerevisiae.G3 (Bethesda). 2022 Jul 29;12(8):jkac157. doi: 10.1093/g3journal/jkac157. G3 (Bethesda). 2022. PMID: 35736360 Free PMC article.
-
Physical links between the nuclear envelope protein Mps3, three alternate replication factor C complexes, and a variant histone in Saccharomyces cerevisiae.DNA Cell Biol. 2012 Jun;31(6):917-24. doi: 10.1089/dna.2011.1493. Epub 2012 Jan 25. DNA Cell Biol. 2012. PMID: 22276573 Free PMC article.
-
Cohesin without cohesion: a novel role for Pds5 in Saccharomyces cerevisiae.PLoS One. 2014 Jun 25;9(6):e100470. doi: 10.1371/journal.pone.0100470. eCollection 2014. PLoS One. 2014. PMID: 24963665 Free PMC article.
-
Cohesin codes - interpreting chromatin architecture and the many facets of cohesin function.J Cell Sci. 2013 Jan 1;126(Pt 1):31-41. doi: 10.1242/jcs.116566. J Cell Sci. 2013. PMID: 23516328 Free PMC article. Review.
-
PCNA Loaders and Unloaders-One Ring That Rules Them All.Genes (Basel). 2021 Nov 18;12(11):1812. doi: 10.3390/genes12111812. Genes (Basel). 2021. PMID: 34828416 Free PMC article. Review.
References
-
- Skibbens RV. Mechanisms of sister chromatid pairing. Inter Rev Cell Mol Biol. 2008;269:283–339. - PubMed
-
- Michaelis C, Ciosk R, Nasmyth K. Cohesins: chromosomal proteins that prevent premature separation of sister chromatids. Cell. 1997;91:35–45. - PubMed
-
- Rankin S, Ayad NG, Kirschner MW. Sororin, a substrate of the anaphase-promoting complex, is required for sister chromatid cohesion in vertebrates. Mol Cell. 2005;18:185–200. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

