microRNA profiling in Epstein-Barr virus-associated B-cell lymphoma
- PMID: 21062812
- PMCID: PMC3061055
- DOI: 10.1093/nar/gkq1043
microRNA profiling in Epstein-Barr virus-associated B-cell lymphoma
Abstract
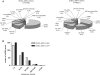
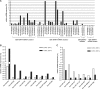
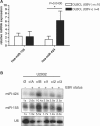
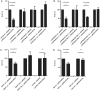
The Epstein-Barr virus (EBV) is an oncogenic human Herpes virus found in ∼15% of diffuse large B-cell lymphoma (DLBCL). EBV encodes miRNAs and induces changes in the cellular miRNA profile of infected cells. MiRNAs are small, non-coding RNAs of ∼19-26 nt which suppress protein synthesis by inducing translational arrest or mRNA degradation. Here, we report a comprehensive miRNA-profiling study and show that hsa-miR-424, -223, -199a-3p, -199a-5p, -27b, -378, -26b, -23a, -23b were upregulated and hsa-miR-155, -20b, -221, -151-3p, -222, -29b/c, -106a were downregulated more than 2-fold due to EBV-infection of DLBCL. All known EBV miRNAs with the exception of the BHRF1 cluster as well as EBV-miR-BART15 and -20 were present. A computational analysis indicated potential targets such as c-MYB, LATS2, c-SKI and SIAH1. We show that c-MYB is targeted by miR-155 and miR-424, that the tumor suppressor SIAH1 is targeted by miR-424, and that c-SKI is potentially regulated by miR-155. Downregulation of SIAH1 protein in DLBCL was demonstrated by immunohistochemistry. The inhibition of SIAH1 is in line with the notion that EBV impedes various pro-apoptotic pathways during tumorigenesis. The down-modulation of the oncogenic c-MYB protein, although counter-intuitive, might be explained by its tight regulation in developmental processes.
Figures

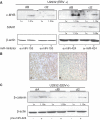
References
-
- Rickinson AB, Kieff E. In: Fields Virology. Knipe DM, Howley PM, editors. Vol. 2. Philadelphia: Lippincott-Raven; 2007. pp. 2655–2700.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

