A novel fluorogenic substrate for the measurement of endothelial lipase activity
- PMID: 21062953
- PMCID: PMC3023558
- DOI: 10.1194/jlr.D007971
A novel fluorogenic substrate for the measurement of endothelial lipase activity
Abstract
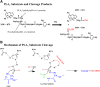
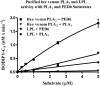
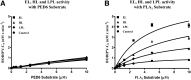
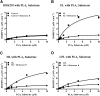
Endothelial lipase (EL) is a phospholipase A1 (PLA1) enzyme that hydrolyzes phospholipids at the sn-1 position to produce lysophospholipids and free fatty acids. Measurement of the PLA1 activity of EL is usually accomplished by the use of substrates that are also hydrolyzed by lipases in other subfamilies such as PLA2 enzymes. In order to distinguish PLA1 activity of EL from PLA2 enzymatic activity in cell-based assays, cell supernatants, and other nonhomogeneous systems, a novel fluorogenic substrate with selectivity toward PLA1 hydrolysis was conceived and characterized. This substrate was preferred by PLA1 enzymes, such as EL and hepatic lipase, and was cleaved with much lower efficiency by lipases that exhibit primarily triglyceride lipase activity, such as LPL or a lipase with PLA2 activity. The phospholipase activity detected by the PLA1 substrate could be inhibited with the small molecule esterase inhibitor ebelactone B. Furthermore, the PLA1 substrate was able to detect EL activity in human umbilical vein endothelial cells in a cell-based assay. This substrate is a useful reagent for identifying modulators of PLA1 enzymes, such as EL, and aiding in characterizing their mechanisms of action.
Figures



Similar articles
-
Structure and function of extracellular phospholipase A1 belonging to the pancreatic lipase gene family.Biochimie. 2007 Feb;89(2):197-204. doi: 10.1016/j.biochi.2006.09.021. Epub 2006 Oct 27. Biochimie. 2007. PMID: 17101204 Review.
-
Measurement of the phospholipase activity of endothelial lipase in mouse plasma.J Lipid Res. 2013 Jan;54(1):282-9. doi: 10.1194/jlr.D031112. Epub 2012 Oct 26. J Lipid Res. 2013. PMID: 23103358 Free PMC article.
-
Current Knowledge on Mammalian Phospholipase A1, Brief History, Structures, Biochemical and Pathophysiological Roles.Molecules. 2022 Apr 12;27(8):2487. doi: 10.3390/molecules27082487. Molecules. 2022. PMID: 35458682 Free PMC article. Review.
-
Cardiolipin hydrolysis by human phospholipases A2. The multiple enzymatic activities of human cytosolic phospholipase A2.Biochim Biophys Acta. 1998 Feb 5;1390(1):65-72. doi: 10.1016/s0005-2760(97)00170-7. Biochim Biophys Acta. 1998. PMID: 9487141
-
Selective measurement of NAPE-PLD activity via a PLA1/2-resistant fluorogenic N-acyl-phosphatidylethanolamine analog.J Lipid Res. 2022 Jan;63(1):100156. doi: 10.1016/j.jlr.2021.100156. Epub 2021 Nov 26. J Lipid Res. 2022. PMID: 34843683 Free PMC article.
Cited by
-
Sialylation of Asparagine 612 Inhibits Aconitase Activity during Mouse Sperm Capacitation; a Possible Mechanism for the Switch from Oxidative Phosphorylation to Glycolysis.Mol Cell Proteomics. 2020 Nov;19(11):1860-1875. doi: 10.1074/mcp.RA120.002109. Epub 2020 Aug 24. Mol Cell Proteomics. 2020. PMID: 32839225 Free PMC article.
-
Activity and Crystal Structure of the Adherent-Invasive Escherichia coli Tle3/Tli3 T6SS Effector/Immunity Complex Determined Using an AlphaFold2 Predicted Model.Int J Mol Sci. 2023 Jan 16;24(2):1740. doi: 10.3390/ijms24021740. Int J Mol Sci. 2023. PMID: 36675258 Free PMC article.
-
Phospholipase B is activated in response to sterol removal and stimulates acrosome exocytosis in murine sperm.J Biol Chem. 2013 Sep 27;288(39):28104-15. doi: 10.1074/jbc.M113.450981. Epub 2013 Aug 13. J Biol Chem. 2013. PMID: 23943622 Free PMC article.
-
Structural basis for loading and inhibition of a bacterial T6SS phospholipase effector by the VgrG spike.EMBO J. 2020 Jun 2;39(11):e104129. doi: 10.15252/embj.2019104129. Epub 2020 Apr 30. EMBO J. 2020. PMID: 32350888 Free PMC article.
-
Cannabinol modulates the endocannabinoid system and shows TRPV1-mediated anti-inflammatory properties in human keratinocytes.Biofactors. 2025 Jan-Feb;51(1):e2122. doi: 10.1002/biof.2122. Epub 2024 Sep 14. Biofactors. 2025. PMID: 39275884 Free PMC article.
References
-
- Brown R. J., Rader D. J. 2007. Lipases as modulators of atherosclerosis in murine models. Curr. Drug Targets. 8: 1307–1319. - PubMed
-
- Duffy D., Rader D. J. 2009. Update on strategies to increase HDL quantity and function. Nat. Rev. Cardiol. 6: 455–463. - PubMed
-
- Choi S. Y., Hirata K., Ishida T., Quertermous T., Cooper A. D. 2002. Endothelial lipase: a new lipase on the block. J. Lipid Res. 43: 1763–1769. - PubMed
-
- Duong M., Psaltis M., Rader D. J., Marchadier D., Barter P. J., Rye K. A. 2003. Evidence that hepatic lipase and endothelial lipase have different substrate specificities for high-density lipoprotein phospholipids. Biochemistry. 42: 13778–13785. - PubMed
-
- McCoy M. G., Sun G. S., Marchadier D., Maugeais C., Glick J. M., Rader D. J. 2002. Characterization of the lipolytic activity of endothelial lipase. J. Lipid Res. 43: 921–929. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

