CAMP (C13orf8, ZNF828) is a novel regulator of kinetochore-microtubule attachment
- PMID: 21063390
- PMCID: PMC3020106
- DOI: 10.1038/emboj.2010.276
CAMP (C13orf8, ZNF828) is a novel regulator of kinetochore-microtubule attachment
Abstract
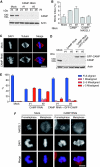
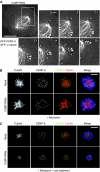
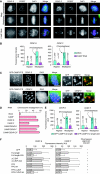
Proper attachment of microtubules to kinetochores is essential for accurate chromosome segregation. Here, we report a novel protein involved in kinetochore-microtubule attachment, chromosome alignment-maintaining phosphoprotein (CAMP) (C13orf8, ZNF828). CAMP is a zinc-finger protein containing three characteristic repeat motifs termed the WK, SPE, and FPE motifs. CAMP localizes to chromosomes and the spindle including kinetochores, and undergoes CDK1-dependent phosphorylation at multiple sites during mitosis. CAMP-depleted cells showed severe chromosome misalignment, which was associated with the poor resistance of K-fibres to the tension exerted upon establishment of sister kinetochore bi-orientation. We found that the FPE region, which is responsible for spindle and kinetochore localization, is essential for proper chromosome alignment. The C-terminal region containing the zinc-finger domains negatively regulates chromosome alignment, and phosphorylation in the FPE region counteracts this regulation. Kinetochore localization of CENP-E and CENP-F was affected by CAMP depletion, and by expressing CAMP mutants that cannot functionally rescue CAMP depletion, placing CENP-E and CENP-F as downstream effectors of CAMP. These data suggest that CAMP is required for maintaining kinetochore-microtubule attachment during bi-orientation.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures





Similar articles
-
Septin 7 interacts with centromere-associated protein E and is required for its kinetochore localization.J Biol Chem. 2008 Jul 4;283(27):18916-25. doi: 10.1074/jbc.M710591200. Epub 2008 May 6. J Biol Chem. 2008. PMID: 18460473 Free PMC article.
-
CENP-E kinesin interacts with SKAP protein to orchestrate accurate chromosome segregation in mitosis.J Biol Chem. 2012 Jan 6;287(2):1500-9. doi: 10.1074/jbc.M111.277194. Epub 2011 Nov 22. J Biol Chem. 2012. PMID: 22110139 Free PMC article.
-
BubR1 and CENP-E have antagonistic effects upon the stability of microtubule-kinetochore attachments in Drosophila S2 cell mitosis.Cell Cycle. 2007 Jun 1;6(11):1367-78. doi: 10.4161/cc.6.11.4271. Epub 2007 Jun 11. Cell Cycle. 2007. PMID: 17525528
-
Mechanisms of kinesin-7 CENP-E in kinetochore-microtubule capture and chromosome alignment during cell division.Biol Cell. 2019 Jun;111(6):143-160. doi: 10.1111/boc.201800082. Epub 2019 Feb 26. Biol Cell. 2019. PMID: 30784092 Review.
-
Leaving no-one behind: how CENP-E facilitates chromosome alignment.Essays Biochem. 2020 Sep 4;64(2):313-324. doi: 10.1042/EBC20190073. Essays Biochem. 2020. PMID: 32347304 Free PMC article. Review.
Cited by
-
Deficiency of CHAMP1, a gene related to intellectual disability, causes impaired neuronal development and a mild behavioural phenotype.Brain Commun. 2022 Aug 30;4(5):fcac220. doi: 10.1093/braincomms/fcac220. eCollection 2022. Brain Commun. 2022. PMID: 36106092 Free PMC article.
-
Chromosome alignment-maintaining phosphoprotein CHAMP1 plays a role in cell survival through regulating Mcl-1 expression.Cancer Sci. 2021 Sep;112(9):3711-3721. doi: 10.1111/cas.15018. Epub 2021 Jul 16. Cancer Sci. 2021. PMID: 34107118 Free PMC article.
-
De Novo Mutations in CHAMP1 Cause Intellectual Disability with Severe Speech Impairment.Am J Hum Genet. 2015 Sep 3;97(3):493-500. doi: 10.1016/j.ajhg.2015.08.003. Am J Hum Genet. 2015. PMID: 26340335 Free PMC article.
-
REV7: a small but mighty regulator of genome maintenance and cancer development.Front Oncol. 2025 Jan 7;14:1516165. doi: 10.3389/fonc.2024.1516165. eCollection 2024. Front Oncol. 2025. PMID: 39839778 Free PMC article. Review.
-
[Autosomal dominant intellectual disability-40 caused by a de novo mutation of the CHAMP1 gene: a case report].Zhongguo Dang Dai Er Ke Za Zhi. 2020 Oct;22(10):1131-1134. doi: 10.7499/j.issn.1008-8830.2004163. Zhongguo Dang Dai Er Ke Za Zhi. 2020. PMID: 33059813 Free PMC article. Chinese.
References
-
- Ashar HR, James L, Gray K, Carr D, Black S, Armstrong L, Bishop WR, Kirschmeier P (2000) Farnesyl transferase inhibitors block the farnesylation of CENP-E and CENP-F and alter the association of CENP-E with the microtubules. J Biol Chem 275: 30451–30457 - PubMed
-
- Cahill DP, da Costa LT, Carson-Walter EB, Kinzler KW, Vogelstein B, Lengauer C (1999) Characterization of MAD2B and other mitotic spindle checkpoint genes. Genomics 58: 181–187 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

