Transgenic mice: beyond the knockout
- PMID: 21068085
- PMCID: PMC3044000
- DOI: 10.1152/ajprenal.00082.2010
Transgenic mice: beyond the knockout
Abstract
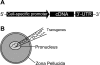
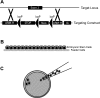
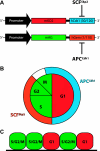
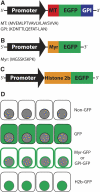
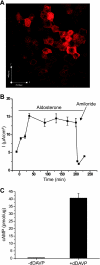
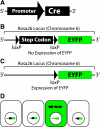
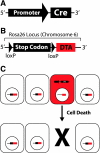
Transgenic mice have had a tremendous impact on biomedical research. Most researchers are familiar with transgenic mice that carry Cre recombinase (Cre) and how they are used to create conditional knockouts. However, some researchers are less familiar with many of the other types of transgenic mice and their applications. For example, transgenic mice can be used to study biochemical and molecular pathways in primary cultures and cell suspensions derived from transgenic mice, cell-cell interactions using multiple fluorescent proteins in the same mouse, and the cell cycle in real time and in the whole animal, and they can be used to perform deep tissue imaging in the whole animal, follow cell lineage during development and disease, and isolate large quantities of a pure cell type directly from organs. These novel transgenic mice and their applications provide the means for studying of molecular and biochemical events in the whole animal that was previously limited to cell cultures. In conclusion, transgenic mice are not just for generating knockouts.
Figures



Similar articles
-
Identification of undescribed Relb expression domains in the murine brain by new Relb:cre-katushka reporter mice.Dev Dyn. 2020 Aug;249(8):983-997. doi: 10.1002/dvdy.170. Epub 2020 May 30. Dev Dyn. 2020. PMID: 32145043
-
A novel reporter mouse strain that expresses enhanced green fluorescent protein upon Cre-mediated recombination.FEBS Lett. 2000 Mar 31;470(3):263-8. doi: 10.1016/s0014-5793(00)01338-7. FEBS Lett. 2000. PMID: 10745079
-
Temporally controlled site-specific mutagenesis in the germ cell lineage of the mouse testis.Biol Reprod. 2003 Feb;68(2):553-9. doi: 10.1095/biolreprod.102.005801. Biol Reprod. 2003. PMID: 12533419
-
Cre Activated and Inactivated Recombinant Adeno-Associated Viral Vectors for Neuronal Anatomical Tracing or Activity Manipulation.Curr Protoc Neurosci. 2015 Jul 1;72:1.24.1-1.24.15. doi: 10.1002/0471142301.ns0124s72. Curr Protoc Neurosci. 2015. PMID: 26131660 Free PMC article. Review.
-
In Vivo Genetic Strategies for the Specific Lineage Tracing of Stem Cells.Curr Stem Cell Res Ther. 2019;14(3):230-238. doi: 10.2174/1574888X13666180726110138. Curr Stem Cell Res Ther. 2019. PMID: 30047336 Review.
Cited by
-
Zebrafish: A Resourceful Vertebrate Model to Investigate Skeletal Disorders.Front Endocrinol (Lausanne). 2020 Jul 31;11:489. doi: 10.3389/fendo.2020.00489. eCollection 2020. Front Endocrinol (Lausanne). 2020. PMID: 32849280 Free PMC article. Review.
-
A versatile toolbox for semi-automatic cell-by-cell object-based colocalization analysis.Sci Rep. 2020 Nov 4;10(1):19027. doi: 10.1038/s41598-020-75835-7. Sci Rep. 2020. PMID: 33149236 Free PMC article.
-
Using transgenic reporters to visualize bone and cartilage signaling during development in vivo.Front Endocrinol (Lausanne). 2012 Jul 18;3:91. doi: 10.3389/fendo.2012.00091. eCollection 2012. Front Endocrinol (Lausanne). 2012. PMID: 22826703 Free PMC article.
-
Replication-competent influenza A viruses expressing a red fluorescent protein.Virology. 2015 Feb;476:206-216. doi: 10.1016/j.virol.2014.12.006. Epub 2014 Dec 30. Virology. 2015. PMID: 25553516 Free PMC article.
-
Incomplete cre-mediated excision leads to phenotypic differences between Stra8-iCre; Mov10l1(lox/lox) and Stra8-iCre; Mov10l1(lox/Δ) mice.Genesis. 2013 Jul;51(7):481-90. doi: 10.1002/dvg.22389. Epub 2013 Mar 30. Genesis. 2013. PMID: 23554062 Free PMC article.
References
-
- Amoh Y, Katsuoka K, Hoffman RM. Color-coded fluorescent protein imaging of angiogenesis: the AngioMouse models. Curr Pharm Des 14: 3810–3819, 2008 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

