Herpes simplex virus type 1 helicase-primase: DNA binding and consequent protein oligomerization and primase activation
- PMID: 21068246
- PMCID: PMC3019990
- DOI: 10.1128/JVI.01690-10
Herpes simplex virus type 1 helicase-primase: DNA binding and consequent protein oligomerization and primase activation
Abstract
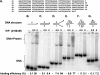
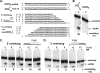
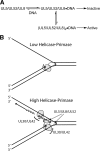
The heterotrimeric helicase-primase complex of herpes simplex virus type I (HSV-1), consisting of UL5, UL8, and UL52, possesses 5' to 3' helicase, single-stranded DNA (ssDNA)-dependent ATPase, primase, and DNA binding activities. In this study we confirm that the UL5-UL8-UL52 complex has higher affinity for forked DNA than for ssDNA and fails to bind to fully annealed double-stranded DNA substrates. In addition, we show that a single-stranded overhang of greater than 6 nucleotides is required for efficient enzyme loading and unwinding. Electrophoretic mobility shift assays and surface plasmon resonance analysis provide additional quantitative information about how the UL5-UL8-UL52 complex associates with the replication fork. Although it has previously been reported that in the absence of DNA and nucleoside triphosphates the UL5-UL8-UL52 complex exists as a monomer in solution, we now present evidence that in the presence of forked DNA and AMP-PNP, higher-order complexes can form. Electrophoretic mobility shift assays reveal two discrete complexes with different mobilities only when helicase-primase is bound to DNA containing a single-stranded region, and surface plasmon resonance analysis confirms larger amounts of the complex bound to forked substrates than to single-overhang substrates. Furthermore, we show that primase activity exhibits a cooperative dependence on protein concentration while ATPase and helicase activities do not. Taken together, these data suggest that the primase activity of the helicase-primase requires formation of a dimer or higher-order structure while ATPase activity does not. Importantly, this provides a simple mechanism for generating a two-polymerase replisome at the replication fork.
Figures







Similar articles
-
The UL5 and UL52 subunits of the herpes simplex virus type 1 helicase-primase subcomplex exhibit a complex interdependence for DNA binding.J Biol Chem. 2001 May 18;276(20):17610-9. doi: 10.1074/jbc.M010107200. Epub 2001 Jan 25. J Biol Chem. 2001. PMID: 11278618
-
Herpes simplex virus-1 helicase-primase: roles of each subunit in DNA binding and phosphodiester bond formation.Biochemistry. 2009 Nov 3;48(43):10199-207. doi: 10.1021/bi9010144. Biochemistry. 2009. PMID: 19788334 Free PMC article.
-
The UL8 subunit of the helicase-primase complex of herpes simplex virus promotes DNA annealing and has a high affinity for replication forks.J Biol Chem. 2017 Sep 22;292(38):15611-15621. doi: 10.1074/jbc.M117.799064. Epub 2017 Jul 25. J Biol Chem. 2017. PMID: 28743747 Free PMC article.
-
The three-component helicase/primase complex of herpes simplex virus-1.Open Biol. 2021 Jun;11(6):210011. doi: 10.1098/rsob.210011. Epub 2021 Jun 9. Open Biol. 2021. PMID: 34102080 Free PMC article. Review.
-
The two helicases of herpes simplex virus type 1 (HSV-1).Front Biosci. 2006 Sep 1;11:2213-23. doi: 10.2741/1964. Front Biosci. 2006. PMID: 16720308 Review.
Cited by
-
CRISPR-Associated Primase-Polymerases are implicated in prokaryotic CRISPR-Cas adaptation.Nat Commun. 2021 Jun 17;12(1):3690. doi: 10.1038/s41467-021-23535-9. Nat Commun. 2021. PMID: 34140468 Free PMC article.
-
Herpes simplex virus 1 ICP8 mutant lacking annealing activity is deficient for viral DNA replication.Proc Natl Acad Sci U S A. 2019 Jan 15;116(3):1033-1042. doi: 10.1073/pnas.1817642116. Epub 2018 Dec 31. Proc Natl Acad Sci U S A. 2019. PMID: 30598436 Free PMC article.
-
Replication and recombination of herpes simplex virus DNA.J Biol Chem. 2011 May 6;286(18):15619-24. doi: 10.1074/jbc.R111.233981. Epub 2011 Mar 1. J Biol Chem. 2011. PMID: 21362621 Free PMC article. Review.
-
Antiviral activity of PHA767491 against human herpes simplex virus in vitro and in vivo.BMC Infect Dis. 2017 Mar 20;17(1):217. doi: 10.1186/s12879-017-2305-0. BMC Infect Dis. 2017. PMID: 28320320 Free PMC article.
-
Protein Displacement by Herpes Helicase-Primase and the Key Role of UL42 during Helicase-Coupled DNA Synthesis by the Herpes Polymerase.Biochemistry. 2017 May 30;56(21):2651-2662. doi: 10.1021/acs.biochem.6b01128. Epub 2017 May 19. Biochemistry. 2017. PMID: 28505413 Free PMC article.
References
-
- Aslani, A., M. Olsson, and P. Elias. 2002. ATP-dependent unwinding of a minimal origin of DNA replication by the origin-binding protein and the single-strand DNA-binding protein ICP8 from herpes simplex virus type I. J. Biol. Chem. 277:41204-41212. - PubMed
-
- Biswas, N., and S. K. Weller. 1999. A mutation in the C-terminal putative Zn2+ finger motif of UL52 severely affects the biochemical activities of the HSV-1 helicase-primase subcomplex. J. Biol. Chem. 274:8068-8076. - PubMed
-
- Biswas, N., and S. K. Weller. 2001. The UL5 and UL52 subunits of the herpes simplex virus type 1 helicase-primase subcomplex exhibit a complex interdependence for DNA binding. J. Biol. Chem. 276:17610-17619. - PubMed
-
- Biswas, S., and H. J. Field. 2008. Herpes simplex virus helicase-primase inhibitors: recent findings from the study of drug resistance mutations. Antivir. Chem. Chemother. 19:1-6. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

