Targets of somatic hypermutation within immunoglobulin light chain genes in zebrafish
- PMID: 21070232
- PMCID: PMC3050447
- DOI: 10.1111/j.1365-2567.2010.03358.x
Targets of somatic hypermutation within immunoglobulin light chain genes in zebrafish
Abstract
In mammals, somatic hypermutation (SHM) of immunoglobulin (Ig) genes is critical for the generation of high-affinity antibodies and effective immune responses. Knowledge of sequence-specific biases in the targeting of somatic mutations can be useful for studies aimed at understanding antibody repertoires produced in response to infections, B-cell neoplasms, or autoimmune disease. To evaluate potential nucleotide targets of somatic mutation in zebrafish (Danio rerio), an enriched IgL cDNA library was constructed and > 250 randomly selected clones were sequenced and analysed. In total, 55 unique VJ-C sequences were identified encoding a total of 125 mutations. Mutations were most prevalent in V(L) with a bias towards single base transitions and increased mutation in the complementarity-determining regions (CDRs). Overall, mutations were overrepresented at WRCH/DGYW motifs suggestive of activation-induced cytidine deaminase (AID) targeting which is common in mice and humans. In contrast to mammalian models, N and P addition was not observed and mutations at AID hotspots were largely restricted to palindromic WRCH/DGYW motifs. Mutability indexes for di- and trinucleotide combinations confirmed C/G targets within WRCH/DGYW motifs to be statistically significant mutational hotspots and showed trinucleotides ATC and ATG to be mutation coldspots. Additive mutations in VJ-C sequences revealed patterns of clonal expansion consistent with affinity maturation responses seen in higher vertebrates. Taken together, the data reveal specific nucleotide targets of SHM in zebrafish and suggest that AID and affinity maturation contribute to antibody diversification in this emerging immunological model.
Figures



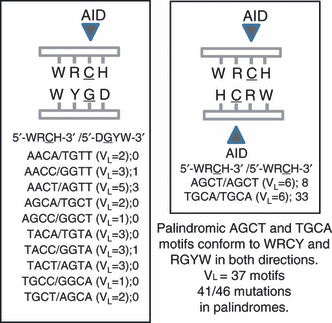
References
-
- Weigert MG, Cesari IM, Yonkovich SJ, Cohn M. Variability in the lambda light chain sequences of mouse antibody. Nature. 1970;228:1045–7. - PubMed
-
- Harris RS, Kong Q, Maizels N. Somatic hypermutation and the three R’s: repair, replication and recombination. Mutat Res. 1999;436:157–78. - PubMed
-
- Maizels N, Scharff MD. Molecular mechanisms of hypermutation. In: Neuberger M, Honjo T, Alt FW, editors. Molecular Biology of B Cells. New York: Academic Press; 2004. pp. 327–38.
-
- Danilova N, Bussmann J, Jekosch K, Steiner LA. The immunoglobulin heavy-chain locus in zebrafish: identification and expression of a previously unknown isotype, immunoglobulin Z. Nat Immunol. 2005;6:295–302. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

