Two LysM receptor molecules, CEBiP and OsCERK1, cooperatively regulate chitin elicitor signaling in rice
- PMID: 21070404
- PMCID: PMC2996852
- DOI: 10.1111/j.1365-313X.2010.04324.x
Two LysM receptor molecules, CEBiP and OsCERK1, cooperatively regulate chitin elicitor signaling in rice
Abstract
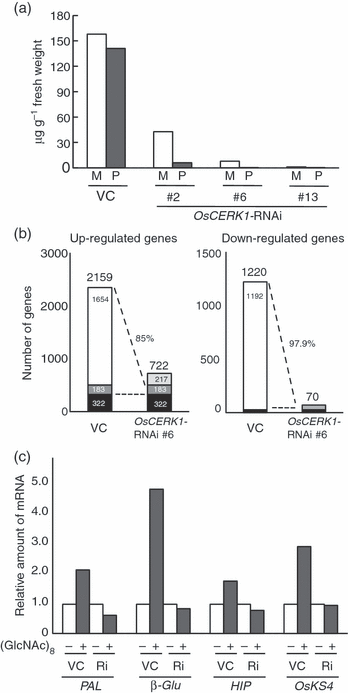
Chitin is a major molecular pattern for various fungi, and its fragments, chitin oligosaccharides, are known to induce various defense responses in plant cells. A plasma membrane glycoprotein, CEBiP (chitin elicitor binding protein) and a receptor kinase, CERK1 (chitin elicitor receptor kinase) (also known as LysM-RLK1), were identified as critical components for chitin signaling in rice and Arabidopsis, respectively. However, it is not known whether each plant species requires both of these two types of molecules for chitin signaling, nor the relationships between these molecules in membrane signaling. We report here that rice cells require a LysM receptor-like kinase, OsCERK1, in addition to CEBiP, for chitin signaling. Knockdown of OsCERK1 resulted in marked suppression of the defense responses induced by chitin oligosaccharides, indicating that OsCERK1 is essential for chitin signaling in rice. The results of a yeast two-hybrid assay indicated that both CEBiP and OsCERK1 have the potential to form hetero- or homo-oligomers. Immunoprecipitation using a membrane preparation from rice cells treated with chitin oligosaccharides suggested the ligand-induced formation of a receptor complex containing both CEBiP and OsCERK1. Blue native PAGE and chemical cross-linking experiments also suggested that a major portion of CEBiP exists as homo-oligomers even in the absence of chitin oligosaccharides.
© 2010 The Authors. Journal compilation © 2010 Blackwell Publishing Ltd.
Figures



References
-
- Chinchilla D, Zipfel C, Robatzek S, Kemmerling B, Nurnberger T, Jones JD, Felix G, Boller T. A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature. 2007;448:497–500. - PubMed
-
- Chisholm ST, Coaker G, Day B, Staskawicz BJ. Host–microbe interactions: shaping the evolution of the plant immune response. Cell. 2006;124:803–814. - PubMed
-
- Desaki Y, Miya A, Venkatesh B, Tsuyumu S, Yamane H, Kaku H, Minami E, Shibuya N. Bacterial lipopolysaccharides induce defense responses associated with programmed cell death in rice cells. Plant Cell Physiol. 2006;47:1530–1540. - PubMed
-
- Gimenez-Ibanez S, Hann DR, Ntoukakis V, Petutschnig E, Lipka V, Rathjen JP. AvrPtoB targets the LysM receptor kinase CERK1 to promote bacterial virulence on plants. Curr. Biol. 2009;19:423–429. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

