Are substitution rates and RNA editing correlated?
- PMID: 21070620
- PMCID: PMC2989974
- DOI: 10.1186/1471-2148-10-349
Are substitution rates and RNA editing correlated?
Abstract
Background: RNA editing is a post-transcriptional process that, in seed plants, involves a cytosine to uracil change in messenger RNA, causing the translated protein to differ from that predicted by the DNA sequence. RNA editing occurs extensively in plant mitochondria, but large differences in editing frequencies are found in some groups. The underlying processes responsible for the distribution of edited sites are largely unknown, but gene function, substitution rate, and gene conversion have been proposed to influence editing frequencies.
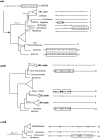
Results: We studied five mitochondrial genes in the monocot order Alismatales, all showing marked differences in editing frequencies among taxa. A general tendency to lose edited sites was observed in all taxa, but this tendency was particularly strong in two clades, with most of the edited sites lost in parallel in two different areas of the phylogeny. This pattern is observed in at least four of the five genes analyzed. Except in the groups that show an unusually low editing frequency, the rate of C-to-T changes in edited sites was not significantly higher that in non-edited 3rd codon positions. This may indicate that selection is not actively removing edited sites in nine of the 12 families of the core Alismatales. In all genes but ccmB, a significant correlation was found between frequency of change in edited sites and synonymous substitution rate. In general, taxa with higher substitution rates tend to have fewer edited sites, as indicated by the phylogenetically independent correlation analyses. The elimination of edited sites in groups that lack or have reduced levels of editing could be a result of gene conversion involving a cDNA copy (retroprocessing). If so, this phenomenon could be relatively common in the Alismatales, and may have affected some groups recurrently. Indirect evidence of retroprocessing without a necessary correlation with substitution rate was found mostly in families Alismataceae and Hydrocharitaceae (e.g., groups that suffered a rapid elimination of all their edited sites, without a change in substitution rate).
Conclusions: The effects of substitution rate, selection, and/or gene conversion on the dynamics of edited sites in plant mitochondria remain poorly understood. Although we found an inverse correlation between substitution rate and editing frequency, this correlation is partially obscured by gene retroprocessing in lineages that have lost most of their edited sites. The presence of processed paralogs in plant mitochondria deserves further study, since most evidence of their occurrence is circumstantial.
Figures


Similar articles
-
Towards a comprehensive picture of C-to-U RNA editing sites in angiosperm mitochondria.Plant Mol Biol. 2018 Jun;97(3):215-231. doi: 10.1007/s11103-018-0734-9. Epub 2018 May 14. Plant Mol Biol. 2018. PMID: 29761268
-
Localized Retroprocessing as a Model of Intron Loss in the Plant Mitochondrial Genome.Genome Biol Evol. 2016 Aug 3;8(7):2176-89. doi: 10.1093/gbe/evw148. Genome Biol Evol. 2016. PMID: 27435795 Free PMC article.
-
Extensive loss of RNA editing sites in rapidly evolving Silene mitochondrial genomes: selection vs. retroprocessing as the driving force.Genetics. 2010 Aug;185(4):1369-80. doi: 10.1534/genetics.110.118000. Epub 2010 May 17. Genetics. 2010. PMID: 20479143 Free PMC article.
-
RNA editing in higher plant mitochondria: analysis of biochemistry and specificity.Biochimie. 1995;77(1-2):79-86. doi: 10.1016/0300-9084(96)88108-9. Biochimie. 1995. PMID: 7599280 Review.
-
RNA editing in plants and its evolution.Annu Rev Genet. 2013;47:335-52. doi: 10.1146/annurev-genet-111212-133519. Annu Rev Genet. 2013. PMID: 24274753 Review.
Cited by
-
High Level of Conservation of Mitochondrial RNA Editing Sites Among Four Populus Species.G3 (Bethesda). 2019 Mar 7;9(3):709-717. doi: 10.1534/g3.118.200763. G3 (Bethesda). 2019. PMID: 30617214 Free PMC article.
-
Chloroplast genomic comparison provides insights into the evolution of seagrasses.BMC Plant Biol. 2023 Feb 22;23(1):104. doi: 10.1186/s12870-023-04119-9. BMC Plant Biol. 2023. PMID: 36814193 Free PMC article.
-
Complete sequencing of the mitochondrial genome of tea plant Camellia sinensis cv. 'Baihaozao': multichromosomal structure, phylogenetic relationships, and adaptive evolutionary analysis.Front Plant Sci. 2025 Jun 13;16:1604404. doi: 10.3389/fpls.2025.1604404. eCollection 2025. Front Plant Sci. 2025. PMID: 40584852 Free PMC article.
-
Towards a comprehensive picture of C-to-U RNA editing sites in angiosperm mitochondria.Plant Mol Biol. 2018 Jun;97(3):215-231. doi: 10.1007/s11103-018-0734-9. Epub 2018 May 14. Plant Mol Biol. 2018. PMID: 29761268
-
Mitochondrial Retroprocessing Promoted Functional Transfers of rpl5 to the Nucleus in Grasses.Mol Biol Evol. 2017 Sep 1;34(9):2340-2354. doi: 10.1093/molbev/msx170. Mol Biol Evol. 2017. PMID: 28541477 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

