The BioGRID Interaction Database: 2011 update
- PMID: 21071413
- PMCID: PMC3013707
- DOI: 10.1093/nar/gkq1116
The BioGRID Interaction Database: 2011 update
Abstract
The Biological General Repository for Interaction Datasets (BioGRID) is a public database that archives and disseminates genetic and protein interaction data from model organisms and humans (http://www.thebiogrid.org). BioGRID currently holds 347,966 interactions (170,162 genetic, 177,804 protein) curated from both high-throughput data sets and individual focused studies, as derived from over 23,000 publications in the primary literature. Complete coverage of the entire literature is maintained for budding yeast (Saccharomyces cerevisiae), fission yeast (Schizosaccharomyces pombe) and thale cress (Arabidopsis thaliana), and efforts to expand curation across multiple metazoan species are underway. The BioGRID houses 48,831 human protein interactions that have been curated from 10,247 publications. Current curation drives are focused on particular areas of biology to enable insights into conserved networks and pathways that are relevant to human health. The BioGRID 3.0 web interface contains new search and display features that enable rapid queries across multiple data types and sources. An automated Interaction Management System (IMS) is used to prioritize, coordinate and track curation across international sites and projects. BioGRID provides interaction data to several model organism databases, resources such as Entrez-Gene and other interaction meta-databases. The entire BioGRID 3.0 data collection may be downloaded in multiple file formats, including PSI MI XML. Source code for BioGRID 3.0 is freely available without any restrictions.
Figures
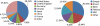

References
-
- Bork P, Jensen LJ, von Mering C, Ramani AK, Lee I, Marcotte EM. Protein interaction networks from yeast to human. Curr. Opin. Struct. Biol. 2004;14:292–299. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

