Shaping the response: the role of FcεRI and Syk expression levels in mast cell signaling
- PMID: 21073233
- PMCID: PMC3057513
- DOI: 10.1049/iet-syb.2010.0006
Shaping the response: the role of FcεRI and Syk expression levels in mast cell signaling
Abstract
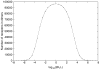
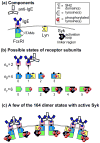
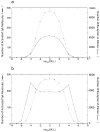
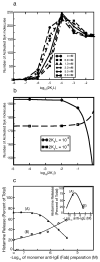
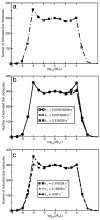
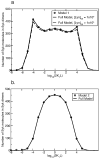
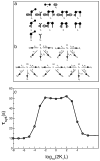
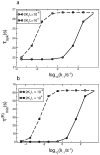
Many receptor systems initiate cell signaling through ligand-induced receptor aggregation. For bivalent ligands binding to mono- or bivalent receptors, a plot of the equilibrium concentration of receptors in aggregates against the log of the free ligand concentration, the cross-linking curve, is symmetric and bell shaped. However, steady state cellular responses initiated through receptor cross-linking may have a different dependence on ligand concentration than the aggregated receptors that initiate and maintain these responses. The authors illustrate by considering the activation of the protein kinase Syk that rapidly occurs after high affinity receptors for IgE, FcεRI, are aggregated on the surface of mast cells and basophils. Using a mathematical model of Syk activation the authors investigate two effects, one straightforward and one less so, that result in Syk activation not qualitatively following the cross-linking curve. Model predictions show that if the mechanism by which Syk is fully activated involves the transphosphorylation of Syk by Syk, then Syk activation curves can be either bell shaped or double humped, depending on the cellular concentrations of Syk and FcεRI. The model also predicts that the Syk activation curve can be non-symmetric with respect to the ligand concentration. The cell can exhibit differential Syk activation at two different ligand concentrations that produce identical distributions of receptor aggregates that form and dissociate at the same rates. The authors discuss how, even though it is only receptor aggregates that trigger responses, differences in total ligand concentration can lead to subtle kinetic effects that yield qualitative differences in the levels of Syk activation.
Figures

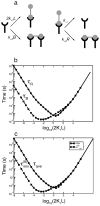
References
-
- Rivera J, Gilfillan AM. Molecular regulation of mast cell activation. J Allergy Clin Immunol. 2006;117:1214–1225. - PubMed
-
- MacGlashan D., Jr IgE and FcεRI regulation. Ann N Y Acad Sci. 2005;1050:73–88. - PubMed
-
- Blank U, Ra C, Miller L, White K, Metzger H, Kinet JP. Complete structure and expression in transfected cells of high affinity IgE receptor. Nature. 1989;337:187–189. - PubMed
-
- Faeder JR, Hlavacek WS, Reischl I, Blinov ML, Metzger H, Redondo A, et al. Investigation of Early Events in FcεRI-Mediated Signaling Using a Detailed Mathematical Model. J Immunol. 2003;170:3769–3781. - PubMed
-
- Kambayashi T, Koretzky GA. Proximal signaling events in FcεRI-mediated mast cell activation. J Allergy Clin Immunol. 2007;119:544–552. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

