Update of the FANTOM web resource: from mammalian transcriptional landscape to its dynamic regulation
- PMID: 21075797
- PMCID: PMC3013704
- DOI: 10.1093/nar/gkq1112
Update of the FANTOM web resource: from mammalian transcriptional landscape to its dynamic regulation
Abstract
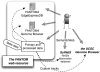
The international Functional Annotation Of the Mammalian Genomes 4 (FANTOM4) research collaboration set out to better understand the transcriptional network that regulates macrophage differentiation and to uncover novel components of the transcriptome employing a series of high-throughput experiments. The primary and unique technique is cap analysis of gene expression (CAGE), sequencing mRNA 5'-ends with a second-generation sequencer to quantify promoter activities even in the absence of gene annotation. Additional genome-wide experiments complement the setup including short RNA sequencing, microarray gene expression profiling on large-scale perturbation experiments and ChIP-chip for epigenetic marks and transcription factors. All the experiments are performed in a differentiation time course of the THP-1 human leukemic cell line. Furthermore, we performed a large-scale mammalian two-hybrid (M2H) assay between transcription factors and monitored their expression profile across human and mouse tissues with qRT-PCR to address combinatorial effects of regulation by transcription factors. These interdependent data have been analyzed individually and in combination with each other and are published in related but distinct papers. We provide all data together with systematic annotation in an integrated view as resource for the scientific community (http://fantom.gsc.riken.jp/4/). Additionally, we assembled a rich set of derived analysis results including published predicted and validated regulatory interactions. Here we introduce the resource and its update after the initial release.
Figures
Similar articles
-
The FANTOM web resource: from mammalian transcriptional landscape to its dynamic regulation.Genome Biol. 2009;10(4):R40. doi: 10.1186/gb-2009-10-4-r40. Epub 2009 Apr 19. Genome Biol. 2009. PMID: 19374775 Free PMC article.
-
Update of the FANTOM web resource: expansion to provide additional transcriptome atlases.Nucleic Acids Res. 2019 Jan 8;47(D1):D752-D758. doi: 10.1093/nar/gky1099. Nucleic Acids Res. 2019. PMID: 30407557 Free PMC article.
-
Macrophages.com: an on-line community resource for innate immunity research.Immunobiology. 2011 Nov;216(11):1203-11. doi: 10.1016/j.imbio.2011.07.025. Epub 2011 Jul 23. Immunobiology. 2011. PMID: 21924793
-
Integrated analysis of the genome and the transcriptome by FANTOM.Brief Bioinform. 2004 Sep;5(3):249-58. doi: 10.1093/bib/5.3.249. Brief Bioinform. 2004. PMID: 15383211 Review.
-
A transcriptional perspective on human macrophage biology.Semin Immunol. 2015 Feb;27(1):44-50. doi: 10.1016/j.smim.2015.02.001. Epub 2015 Apr 3. Semin Immunol. 2015. PMID: 25843246 Review.
Cited by
-
The 24th annual Nucleic Acids Research database issue: a look back and upcoming changes.Nucleic Acids Res. 2017 Jan 4;45(D1):D1-D11. doi: 10.1093/nar/gkw1188. Nucleic Acids Res. 2017. PMID: 28053160 Free PMC article.
-
Strategies to identify long noncoding RNAs involved in gene regulation.Cell Biosci. 2012 Nov 6;2(1):37. doi: 10.1186/2045-3701-2-37. Cell Biosci. 2012. PMID: 23126680 Free PMC article.
-
The plausible reason why the length of 5' untranslated region is unrelated to organismal complexity.BMC Res Notes. 2011 Aug 27;4:312. doi: 10.1186/1756-0500-4-312. BMC Res Notes. 2011. PMID: 21871111 Free PMC article.
-
Update of the FANTOM web resource: high resolution transcriptome of diverse cell types in mammals.Nucleic Acids Res. 2017 Jan 4;45(D1):D737-D743. doi: 10.1093/nar/gkw995. Epub 2016 Oct 27. Nucleic Acids Res. 2017. PMID: 27794045 Free PMC article.
-
Analysis of impact metrics for the Protein Data Bank.Sci Data. 2018 Oct 16;5:180212. doi: 10.1038/sdata.2018.212. Sci Data. 2018. PMID: 30325351 Free PMC article.
References
-
- Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, et al. The transcriptional landscape of the mammalian genome. Science. 2005;309:1559–1563. - PubMed
-
- Katayama S, Tomaru Y, Kasukawa T, Waki K, Nakanishi M, Nakamura M, Nishida H, Yap CC, Suzuki M, Kawai J, et al. Antisense transcription in the mammalian transcriptome. Science. 2005;309:1564–1566. - PubMed
-
- Kawai J, Shinagawa A, Shibata K, Yoshino M, Itoh M, Ishii Y, Arakawa T, Hara A, Fukunishi Y, Konno H, et al. Functional annotation of a full-length mouse cDNA collection. Nature. 2001;409:685–690. - PubMed
-
- Okazaki Y, Furuno M, Kasukawa T, Adachi J, Bono H, Kondo S, Nikaido I, Osato N, Saito R, Suzuki H, et al. Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs. Nature. 2002;420:563–573. - PubMed
-
- Faulkner GJ, Kimura Y, Daub CO, Wani S, Plessy C, Irvine KM, Schroder K, Cloonan N, Steptoe AL, Lassmann T, et al. The regulated retrotransposon transcriptome of mammalian cells. Nat. Genet. 2009;41:563–571. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous