TESTLoc: protein subcellular localization prediction from EST data
- PMID: 21078192
- PMCID: PMC3000424
- DOI: 10.1186/1471-2105-11-563
TESTLoc: protein subcellular localization prediction from EST data
Abstract
Background: The eukaryotic cell has an intricate architecture with compartments and substructures dedicated to particular biological processes. Knowing the subcellular location of proteins not only indicates how bio-processes are organized in different cellular compartments, but also contributes to unravelling the function of individual proteins. Computational localization prediction is possible based on sequence information alone, and has been successfully applied to proteins from virtually all subcellular compartments and all domains of life. However, we realized that current prediction tools do not perform well on partial protein sequences such as those inferred from Expressed Sequence Tag (EST) data, limiting the exploitation of the large and taxonomically most comprehensive body of sequence information from eukaryotes.
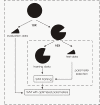
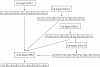
Results: We developed a new predictor, TESTLoc, suited for subcellular localization prediction of proteins based on their partial sequence conceptually translated from ESTs (EST-peptides). Support Vector Machine (SVM) is used as computational method and EST-peptides are represented by different features such as amino acid composition and physicochemical properties. When TESTLoc was applied to the most challenging test case (plant data), it yielded high accuracy (~85%).
Conclusions: TESTLoc is a localization prediction tool tailored for EST data. It provides a variety of models for the users to choose from, and is available for download at http://megasun.bch.umontreal.ca/~shenyq/TESTLoc/TESTLoc.html.
Figures

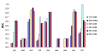
Similar articles
-
'Unite and conquer': enhanced prediction of protein subcellular localization by integrating multiple specialized tools.BMC Bioinformatics. 2007 Oct 29;8:420. doi: 10.1186/1471-2105-8-420. BMC Bioinformatics. 2007. PMID: 17967180 Free PMC article.
-
pSLIP: SVM based protein subcellular localization prediction using multiple physicochemical properties.BMC Bioinformatics. 2005 Jun 17;6:152. doi: 10.1186/1471-2105-6-152. BMC Bioinformatics. 2005. PMID: 15963230 Free PMC article.
-
ESAP plus: a web-based server for EST-SSR marker development.BMC Genomics. 2016 Dec 22;17(Suppl 13):1035. doi: 10.1186/s12864-016-3328-4. BMC Genomics. 2016. PMID: 28155670 Free PMC article.
-
A hitchhiker's guide to expressed sequence tag (EST) analysis.Brief Bioinform. 2007 Jan;8(1):6-21. doi: 10.1093/bib/bbl015. Epub 2006 May 23. Brief Bioinform. 2007. PMID: 16772268 Review.
-
Protein subcellular localization prediction tools.Comput Struct Biotechnol J. 2024 Apr 15;23:1796-1807. doi: 10.1016/j.csbj.2024.04.032. eCollection 2024 Dec. Comput Struct Biotechnol J. 2024. PMID: 38707539 Free PMC article. Review.
Cited by
-
Identification and Expression Pattern of EZH2 in Pig Developing Fetuses.Biomed Res Int. 2020 Oct 5;2020:5315930. doi: 10.1155/2020/5315930. eCollection 2020. Biomed Res Int. 2020. PMID: 33083470 Free PMC article.
-
PSI: a comprehensive and integrative approach for accurate plant subcellular localization prediction.PLoS One. 2013 Oct 23;8(10):e75826. doi: 10.1371/journal.pone.0075826. eCollection 2013. PLoS One. 2013. PMID: 24194827 Free PMC article.
-
Prediction of Protein Subcellular Localization Based on Fusion of Multi-view Features.Molecules. 2019 Mar 6;24(5):919. doi: 10.3390/molecules24050919. Molecules. 2019. PMID: 30845684 Free PMC article.
-
A hybrid distance measure for clustering expressed sequence tags originating from the same gene family.PLoS One. 2012;7(10):e47216. doi: 10.1371/journal.pone.0047216. Epub 2012 Oct 11. PLoS One. 2012. PMID: 23071763 Free PMC article.
-
An ensemble classifier for eukaryotic protein subcellular location prediction using gene ontology categories and amino acid hydrophobicity.PLoS One. 2012;7(1):e31057. doi: 10.1371/journal.pone.0031057. Epub 2012 Jan 30. PLoS One. 2012. PMID: 22303481 Free PMC article.
References
-
- Barbe L, Lundberg E, Oksvold P, Stenius A, Lewin E, Bjorling E, Asplund A, Ponten F, Brismar H, Uhlen M. et al.Toward a confocal subcellular atlas of the human proteome. Mol Cell Proteomics. 2008;7(3):499–508. - PubMed
-
- Yuan HM, Li KL, Ni RJ, Guo WD, Shen Z, Yang CP, Wang BC, Liu GF, Guo CH, Jiang J. A systemic proteomic analysis of Populus chloroplast by using shotgun method. Mol Biol Rep. 2010. in press . - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

