Ancestral and derived protein import pathways in the mitochondrion of Reclinomonas americana
- PMID: 21081480
- PMCID: PMC3080133
- DOI: 10.1093/molbev/msq305
Ancestral and derived protein import pathways in the mitochondrion of Reclinomonas americana
Abstract
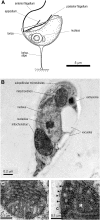
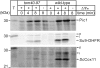
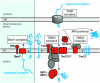
The evolution of mitochondria from ancestral bacteria required that new protein transport machinery be established. Recent controversy over the evolution of these new molecular machines hinges on the degree to which ancestral bacterial transporters contributed during the establishment of the new protein import pathway. Reclinomonas americana is a unicellular eukaryote with the most gene-rich mitochondrial genome known, and the large collection of membrane proteins encoded on the mitochondrial genome of R. americana includes a bacterial-type SecY protein transporter. Analysis of expressed sequence tags shows R. americana also has components of a mitochondrial protein translocase or "translocase in the inner mitochondrial membrane complex." Along with several other membrane proteins encoded on the mitochondrial genome Cox11, an assembly factor for cytochrome c oxidase retains sequence features suggesting that it is assembled by the SecY complex in R. americana. Despite this, protein import studies show that the RaCox11 protein is suited for import into mitochondria and functional complementation if the gene is transferred into the nucleus of yeast. Reclinomonas americana provides direct evidence that bacterial protein transport pathways were retained, alongside the evolving mitochondrial protein import machinery, shedding new light on the process of mitochondrial evolution.
Figures



Similar articles
-
Coupling of import and assembly pathways in mitochondrial protein biogenesis.Biol Chem. 2019 Dec 18;401(1):117-129. doi: 10.1515/hsz-2019-0310. Biol Chem. 2019. PMID: 31513529 Review.
-
A 5 S rRNA gene is present in the mitochondrial genome of the protist Reclinomonas americana but is absent from red algal mitochondrial DNA.J Mol Biol. 1996 Sep 6;261(5):407-13. doi: 10.1006/jmbi.1996.0486. J Mol Biol. 1996. PMID: 8800209
-
A functional Tom70 in the human parasite Blastocystis sp.: implications for the evolution of the mitochondrial import apparatus.Mol Biol Evol. 2011 Jan;28(1):781-91. doi: 10.1093/molbev/msq252. Epub 2010 Sep 24. Mol Biol Evol. 2011. PMID: 20871025
-
Identification of Tim40 that mediates protein sorting to the mitochondrial intermembrane space.J Biol Chem. 2004 Nov 12;279(46):47815-21. doi: 10.1074/jbc.M410272200. Epub 2004 Sep 13. J Biol Chem. 2004. PMID: 15364952
-
The mitochondrial import machinery for preproteins.Crit Rev Biochem Mol Biol. 2001;36(3):291-336. doi: 10.1080/20014091074200. Crit Rev Biochem Mol Biol. 2001. PMID: 11450972 Review.
Cited by
-
Analysis of diverse eukaryotes suggests the existence of an ancestral mitochondrial apparatus derived from the bacterial type II secretion system.Nat Commun. 2021 May 19;12(1):2947. doi: 10.1038/s41467-021-23046-7. Nat Commun. 2021. PMID: 34011950 Free PMC article.
-
Novel Hydrogenosomes in the Microaerophilic Jakobid Stygiella incarcerata.Mol Biol Evol. 2016 Sep;33(9):2318-36. doi: 10.1093/molbev/msw103. Epub 2016 Jun 8. Mol Biol Evol. 2016. PMID: 27280585 Free PMC article.
-
Cytochrome c oxidase: evolution of control via nuclear subunit addition.Biochim Biophys Acta. 2012 Apr;1817(4):590-7. doi: 10.1016/j.bbabio.2011.07.007. Epub 2011 Jul 23. Biochim Biophys Acta. 2012. PMID: 21802404 Free PMC article. Review.
-
The preprotein translocase YidC controls respiratory metabolism in Mycobacterium tuberculosis.Sci Rep. 2016 May 11;6:24998. doi: 10.1038/srep24998. Sci Rep. 2016. PMID: 27166092 Free PMC article.
-
Bacterial Type II Secretion System and Its Mitochondrial Counterpart.mBio. 2023 Apr 25;14(2):e0314522. doi: 10.1128/mbio.03145-22. Epub 2023 Mar 27. mBio. 2023. PMID: 36971557 Free PMC article. Review.
References
-
- Adams KL, Palmer JD. Evolution of mitochondrial gene content: gene loss and transfer to the nucleus. Mol Phylogenet Evol. 2003;29:380–395. - PubMed
-
- Alcock F, Clements A, Webb C, Lithgow T. Tinkering inside the organelle. Science. 2010;327:649–650. - PubMed
-
- Andersson SG, Kurland CG. Origins of mitochondria and hydrogenosomes. Curr Opin Microbiol. 1999;2:535–544. - PubMed
-
- Archibald JM, O'Kelly CJ, Doolittle WF. The chaperonin genes of jakobid and jakobid-like flagellates: implications for eukaryotic evolution. Mol Biol Evol. 2002;19:422–431. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

