Effects of site-directed mutagenesis of mglA on motility and swarming of Myxococcus xanthus
- PMID: 21083931
- PMCID: PMC3000849
- DOI: 10.1186/1471-2180-10-295
Effects of site-directed mutagenesis of mglA on motility and swarming of Myxococcus xanthus
Abstract
Background: The mglA gene from the bacterium Myxococcus xanthus encodes a 22kDa protein related to the Ras superfamily of monomeric GTPases. MglA is required for the normal function of A-motility (adventurous), S-motility (social), fruiting body morphogenesis, and sporulation. MglA and its homologs differ from all eukaryotic and other prokaryotic GTPases because they have a threonine (Thr78) in place of the highly conserved aspartate residue of the consensus PM3 (phosphate-magnesium binding) region. To identify residues critical for MglA function or potential protein interactions, and explore the function of Thr78, the phenotypes of 18 mglA mutants were characterized.
Results: Nine mutants, with mutations predicted to alter residues that bind the guanine base or coordinate magnesium, did not produce detectable MglA. As expected, these mutants were mot- dev- because MglA is essential for these processes. Of the remaining nine mutants, seven showed a wild-type distribution pattern for MglA but fell into two categories with regard to function. Five of the seven mutants exhibited mild phenotypes, but two mutants, T78D and P80A, abolished motility and development. The localization pattern of MglA was abolished in two mutants that were mot- spo- and dev-. These two mutants were predicted to alter surface residues at Asp52 and Thr54, which suggests that these residues are critical for proper localization and may define a protein interaction site. Improving the consensus match with Ras at Thr78 abolished function of MglA. Only the conservative serine substitution was tolerated at this position. Merodiploid constructs revealed that a subset of alleles, including mglAD52A, were dominant and also illustrated that changing the balance of MglA and its co-transcribed partner, MglB, affects A-motility.
Conclusion: Our results suggest that GTP binding is critical for stability of MglA because MglA does not accumulate in mutants that cannot bind GTP. The threonine in PM3 of MglA proteins represents a novel modification of the highly conserved GTPase consensus at this position. The requirement for a hydroxyl group at this position may indicate that MglA is subject to modification under certain conditions. Proper localization of MglA is critical for both motility and development and likely involves protein interactions mediated by residues Asp52 and Thr54.
Figures



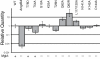


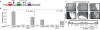


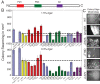
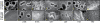

Similar articles
-
Dual specificity of a prokaryotic GTPase-activating protein (GAP) to two small Ras-like GTPases in Myxococcus xanthus.FEBS J. 2021 Mar;288(5):1565-1585. doi: 10.1111/febs.15513. Epub 2020 Sep 3. FEBS J. 2021. PMID: 32772462
-
MglA, a small GTPase, interacts with a tyrosine kinase to control type IV pili-mediated motility and development of Myxococcus xanthus.Mol Microbiol. 2002 Dec;46(5):1399-413. doi: 10.1046/j.1365-2958.2002.03258.x. Mol Microbiol. 2002. PMID: 12453225
-
Function of MglA, a 22-kilodalton protein essential for gliding in Myxococcus xanthus.J Bacteriol. 1991 Dec;173(23):7615-24. doi: 10.1128/jb.173.23.7615-7624.1991. J Bacteriol. 1991. PMID: 1938957 Free PMC article.
-
Polarity of motility systems in Myxococcus xanthus.Curr Opin Microbiol. 2007 Dec;10(6):624-9. doi: 10.1016/j.mib.2007.09.012. Epub 2007 Nov 5. Curr Opin Microbiol. 2007. PMID: 17981496 Free PMC article. Review.
-
Unorthodox regulation of the MglA Ras-like GTPase controlling polarity in Myxococcus xanthus.FEBS Lett. 2023 Mar;597(6):850-864. doi: 10.1002/1873-3468.14565. Epub 2023 Jan 4. FEBS Lett. 2023. PMID: 36520515 Review.
Cited by
-
Evaluation of residue-residue contact prediction in CASP10.Proteins. 2014 Feb;82 Suppl 2(0 2):138-53. doi: 10.1002/prot.24340. Epub 2013 Aug 31. Proteins. 2014. PMID: 23760879 Free PMC article.
-
MasABK proteins interact with proteins of the type IV pilin system to affect social motility of Myxococcus xanthus.PLoS One. 2013;8(1):e54557. doi: 10.1371/journal.pone.0054557. Epub 2013 Jan 16. PLoS One. 2013. PMID: 23342171 Free PMC article.
-
Dual biochemical oscillators may control cellular reversals in Myxococcus xanthus.Biophys J. 2014 Dec 2;107(11):2700-11. doi: 10.1016/j.bpj.2014.09.046. Epub 2014 Dec 2. Biophys J. 2014. PMID: 25468349 Free PMC article.
-
Chemosensory signaling controls motility and subcellular polarity in Myxococcus xanthus.Curr Opin Microbiol. 2012 Dec;15(6):751-7. doi: 10.1016/j.mib.2012.10.005. Epub 2012 Nov 8. Curr Opin Microbiol. 2012. PMID: 23142584 Free PMC article. Review.
-
Ras GTPase-like protein MglA, a controller of bacterial social-motility in Myxobacteria, has evolved to control bacterial predation by Bdellovibrio.PLoS Genet. 2014 Apr 10;10(4):e1004253. doi: 10.1371/journal.pgen.1004253. eCollection 2014 Apr. PLoS Genet. 2014. PMID: 24721965 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

