A new, lineage specific, autoup-regulation mechanism for human glucocorticoid receptor gene expression in 697 pre-B-acute lymphoblastic leukemia cells
- PMID: 21084380
- PMCID: PMC3033051
- DOI: 10.1210/me.2010-0249
A new, lineage specific, autoup-regulation mechanism for human glucocorticoid receptor gene expression in 697 pre-B-acute lymphoblastic leukemia cells
Abstract
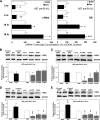
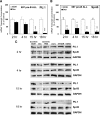
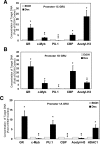
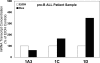
Glucocorticoid (GC) steroid hormones induce apoptosis in acute lymphoblastic leukemia (ALL). Autoup-regulation of human GC receptor (hGR) levels is associated with sensitivity to GC-mediated apoptosis. Among the major hGR promoters expressed in 697 pre-B-ALL cells (1A, 1B, 1C, and 1D), only promoters 1C and 1D are selectively activated by the hormone. Promoter 1B is unresponsive, and promoter 1A is down-regulated by dexamethasone (Dex) in 697 cells, whereas they are both up-regulated in CEM-C7 T-ALL cells. Autoup-regulation of promoter 1C and 1D in 697 cells requires sequences containing GC response units (GRUs) (1C GRU, -2915/-2956; 1D GRU, -4525/-4559) that were identified previously in CEM-C7 cells. These GRUs potentially bind GR, c-myeloblastosis (c-Myb), and E-twenty six (Ets) proteins; 697 cells express high levels of c-Myb protein, as well as the E-twenty six family protein members, PU.1 and Spi-B. Dex treatment in 697 cells elevates the expression of c-Myb and decreases levels of both Spi-B and PU.1. Chromatin immunoprecipitation assays revealed the specific recruitment of GR, c-Myb, and cAMP response element-binding protein binding protein to the 1C and 1D GRUs upon Dex treatment, correlating to observed autoup-regulated activity in these two promoters. These data suggest a hormone activated, lineage-specific mechanism to control the autoup-regulation of hGR gene expression in 697 pre-B-ALL cells via steroid-mediated changes in GR coregulator expression. These findings may be helpful in understanding the mechanism that determines the sensitivity of B-ALL leukemia cells to hormone-induced apoptosis.
Figures




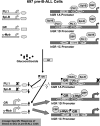
References
-
- Pui CH, Robison LL, Look AT 2008 Acute lymphoblastic leukaemia. Lancet 371:1030–1043 - PubMed
-
- Nanni P, Nicoletti G, Prodi G, Galli MC, De Giovanni C, Grilli S, Lollini PL, Gobbi M, Cavo M, Tura S 1982 Glucocorticoid receptor and in vitro sensitivity to steroid hormones in human lymphoproliferative diseases and myeloid leukemia. Cancer 49:623–632 - PubMed
-
- Gaynon PS, Carrel, AL 1999 Glucocorticosteroid therapy in childhood acute lymphoblastic leukemia. Adv Exp Med Biol 457:593–605 - PubMed
-
- Pui CH, Evans, WE 2006 Treatment of acute lymphoblastic leukemia. N Engl J Med 354:166–178 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

