Functional analysis of a novel motif conserved across geminivirus Rep proteins
- PMID: 21084480
- PMCID: PMC3020519
- DOI: 10.1128/JVI.02143-10
Functional analysis of a novel motif conserved across geminivirus Rep proteins
Abstract
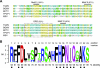
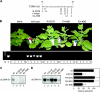
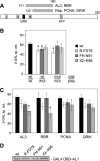
Members of the Geminiviridae have single-stranded DNA genomes that replicate in nuclei of infected plant cells. All geminiviruses encode a conserved protein (Rep) that catalyzes initiation of rolling-circle replication. Earlier studies showed that three conserved motifs-motifs I, II, and III-in the N termini of geminivirus Rep proteins are essential for function. In this study, we identified a fourth sequence, designated GRS (geminivirus Rep sequence), in the Rep N terminus that displays high amino acid sequence conservation across all geminivirus genera. Using the Rep protein of Tomato golden mosaic virus (TGMV AL1), we show that GRS mutants are not infectious in plants and do not support viral genome replication in tobacco protoplasts. GRS mutants are competent for protein-protein interactions and for both double- and single-stranded DNA binding, indicating that the mutations did not impair its global conformation. In contrast, GRS mutants are unable to specifically cleave single-stranded DNA, which is required to initiate rolling-circle replication. Interestingly, the Rep proteins of phytoplasmal and algal plasmids also contain GRS-related sequences. Modeling of the TGMV AL1 N terminus suggested that GRS mutations alter the relative positioning of motif II, which coordinates metal ions, and motif III, which contains the tyrosine involved in DNA cleavage. Together, these results established that the GRS is a conserved, essential motif characteristic of an ancient lineage of rolling-circle initiators and support the idea that geminiviruses may have evolved from plasmids associated with phytoplasma or algae.
Figures


References
-
- Arguello-Astorga, G. R., and R. Ruiz-Medrano. 2001. An iteron-related domain is associated to motif 1 in the replication proteins of geminiviruses: identification of potential interacting amino acid base pairs by a comparative approach. Arch. Virol. 146:1465-1485. - PubMed
-
- Arnold, K., L. Bordoli, J. Kopp, and T. Schwede. 2006. The SWISS-MODEL workspace: a web-based environment for protein structure homology modeling. Bioinformatics 22:195-201. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

