The Mek1 phosphorylation cascade plays a role in meiotic recombination of Schizosaccharomyces pombe
- PMID: 21084840
- PMCID: PMC3048037
- DOI: 10.4161/cc.9.23.14050
The Mek1 phosphorylation cascade plays a role in meiotic recombination of Schizosaccharomyces pombe
Abstract
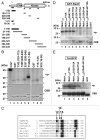
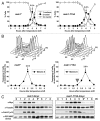
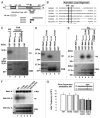
Mek1 is a Chk2/Rad53/Cds1-related protein kinase that is required for proper meiotic progression of Schizosaccharomyces pombe. However, the molecular mechanisms of Mek1 regulation and Mek1 phosphorylation targets are unclear. Here, we report that Mek1 is phosphorylated at serine-12 (S12), S14 and threonine-15 (T15) by Rad3 (ATR) and/or Tel1 (ATM) kinases that are activated by meiotic programmed double-strand breaks (DSBs). Mutations of these sites by alanine replacement caused abnormal meiotic progression and recombination rates. Phosphorylation of these sites triggers autophosphorylation of Mek1; indeed, alanine replacement mutations of Mek1-T318 and -T322 residues in the activation loop of Mek1 reduced Mek1 kinase activity and meiotic recombination rates. Substrates of Mek1 include Mus81-T275, Rdh54-T6 and Rdh54-T673. Mus81-T275 is known to regulate the Mus81 function in DNA cleavage, whereas Rdh54-T6A/T673A mutant cells showed abnormal meiotic recombination. Taken together, we conclude that the phosphorylation of Mek1 by Rad3 or Tel1, Mek1 autophosphorylation and Mus81 or Rdh54 phosphorylation by Mek1 regulate meiotic progression in S. pombe.
Figures




References
-
- Ciccia A, McDonald N, West SC. Structural and Functional Relationships of the XPF/MUS81 Family of Proteins. Annu Rev Biochem. 2008;77:259–287. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
