The peptidoglycan-binding protein FimV promotes assembly of the Pseudomonas aeruginosa type IV pilus secretin
- PMID: 21097635
- PMCID: PMC3019827
- DOI: 10.1128/JB.01048-10
The peptidoglycan-binding protein FimV promotes assembly of the Pseudomonas aeruginosa type IV pilus secretin
Abstract
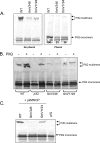
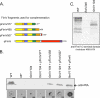
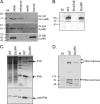
The Pseudomonas aeruginosa inner membrane protein FimV is among several proteins of unknown function required for type IV pilus-mediated twitching motility, arising from extension and retraction of pili from their site of assembly in the inner membrane. The pili transit the periplasm and peptidoglycan (PG) layer, ultimately exiting the cell through the PilQ secretin. Although fimV mutants are nonmotile, they are susceptible to killing by pilus-specific bacteriophage, a hallmark of retractable surface pili. Here we show that levels of recoverable surface pili were markedly decreased in fimV pilT retraction-deficient mutants compared with levels in the pilT control, demonstrating that FimV acts at the level of pilus assembly. Levels of inner membrane assembly subcomplex proteins PilM/N/O/P were decreased in fimV mutants, but supplementation of these components in trans did not restore pilus assembly or motility. Loss of FimV dramatically reduced the levels of the PilQ secretin multimer through which pili exit the cell, in part due to decreased levels of PilQ monomers, while PilF pilotin levels were unchanged. Expression of pilQ in trans in the wild type or fimV mutants increased total PilQ monomer levels but did not alter secretin multimer levels or motility. PG pulldown assays showed that the N terminus of FimV bound PG in a LysM motif-dependent manner, and a mutant with an in-frame chromosomal deletion of the LysM motif had reduced motility, secretin levels, and surface piliation. Together, our data show that FimV's role in pilus assembly is to promote secretin formation and that this function depends upon its PG-binding domain.
Figures


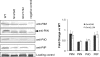
References
-
- Ahn, K. S., U. Ha, J. Jia, D. Wu, and S. Jin. 2004. The truA gene of Pseudomonas aeruginosa is required for the expression of type III secretory genes. Microbiology 150:539-547. - PubMed
-
- Ayers, M., P. L. Howell, and L. L. Burrows. 2010. Architecture of the type II secretion and type IV pilus machineries. Future Microbiol. 5:1203-1218. - PubMed
-
- Ayers, M., et al. 2009. PilM/N/O/P proteins form an inner membrane complex that affects the stability of the Pseudomonas aeruginosa type IV pilus secretin. J. Mol. Biol. 394:128-142. - PubMed
-
- Bateman, A., and M. Bycroft. 2000. The structure of a LysM domain from E. coli membrane-bound lytic murein transglycosylase D (MltD). J. Mol. Biol. 299:1113-1119. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

