RNA polymerase 1-driven transcription as a mediator of BDNF-induced neurite outgrowth
- PMID: 21098478
- PMCID: PMC3039349
- DOI: 10.1074/jbc.M110.170134
RNA polymerase 1-driven transcription as a mediator of BDNF-induced neurite outgrowth
Abstract
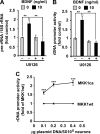
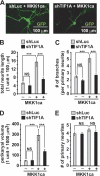
Neurite outgrowth is essential for development of the nervous system. Neurotrophins including BDNF are among extracellular signals that regulate neurite outgrowth. The ERK1/2 pathway contributes to intracellular signaling networks transducing the pro-neuritic effects of BDNF. In the nucleolus, RNA polymerase-1 (Pol1)-mediated transcription regulates ribosomal biogenesis, enabling cellular protein synthesis and growth. Hence, we tested the possibility that Pol1 is an effector for pro-neuritic signals such as BDNF. We report that Pol1-mediated nucleolar transcription was increased by BDNF in an ERK1/2-dependent manner in rat forebrain neurons. Conversely, in cultured hippocampal neurons, knockdown of a Pol1 coactivator, transcription initiation factor 1A (TIF1A), attenuated BDNF- or ERK1/2-induced neurite outgrowth. Also, upon overexpression, a constitutively active mutant of TIF1A strongly promoted neurite outgrowth, including increases in total neurite length and branching. Finally, overexpression of wild-type TIF1A enhanced the pro-neuritic effects of ERK1/2 activation. These observations indicate that the Pol1-mediated nucleolar transcription regulates neurite outgrowth and serves as a major pro-neuritic effector of the BDNF-activated ERK1/2 pathway. Thus, development of the nervous system appears critically dependent on the nucleolus.
Figures




Similar articles
-
New function of the adaptor protein SH2B1 in brain-derived neurotrophic factor-induced neurite outgrowth.PLoS One. 2013 Nov 15;8(11):e79619. doi: 10.1371/journal.pone.0079619. eCollection 2013. PLoS One. 2013. PMID: 24260264 Free PMC article.
-
Role of megakaryoblastic acute leukemia-1 in ERK1/2-dependent stimulation of serum response factor-driven transcription by BDNF or increased synaptic activity.J Neurosci. 2006 Sep 27;26(39):10020-32. doi: 10.1523/JNEUROSCI.2644-06.2006. J Neurosci. 2006. PMID: 17005865 Free PMC article.
-
Dok-4 regulates GDNF-dependent neurite outgrowth through downstream activation of Rap1 and mitogen-activated protein kinase.J Cell Sci. 2006 Aug 1;119(Pt 15):3067-77. doi: 10.1242/jcs.03043. Epub 2006 Jul 4. J Cell Sci. 2006. PMID: 16820412
-
Life on a planet of its own: regulation of RNA polymerase I transcription in the nucleolus.Genes Dev. 2003 Jul 15;17(14):1691-702. doi: 10.1101/gad.1098503R. Genes Dev. 2003. PMID: 12865296 Review. No abstract available.
-
Neurotoxic mechanisms of DNA damage: focus on transcriptional inhibition.J Neurochem. 2010 Sep;114(6):1537-49. doi: 10.1111/j.1471-4159.2010.06859.x. Epub 2010 Aug 19. J Neurochem. 2010. PMID: 20557419 Free PMC article. Review.
Cited by
-
Evidence that vitronectin is a potent migration-enhancing factor for cancer cells chaperoned by fibrinogen: a novel view of the metastasis of cancer cells to low-fibrinogen lymphatics and body cavities.Oncotarget. 2016 Oct 25;7(43):69829-69843. doi: 10.18632/oncotarget.12003. Oncotarget. 2016. PMID: 27634880 Free PMC article.
-
Nucleolar Enrichment of Brain Proteins with Critical Roles in Human Neurodevelopment.Mol Cell Proteomics. 2016 Jun;15(6):2055-75. doi: 10.1074/mcp.M115.051920. Epub 2016 Apr 6. Mol Cell Proteomics. 2016. PMID: 27053602 Free PMC article.
-
A neuroprotective phase precedes striatal degeneration upon nucleolar stress.Cell Death Differ. 2013 Nov;20(11):1455-64. doi: 10.1038/cdd.2013.66. Epub 2013 Jun 14. Cell Death Differ. 2013. PMID: 23764776 Free PMC article.
-
Transcriptional Profiling Reveals Ribosome Biogenesis, Microtubule Dynamics and Expression of Specific lncRNAs to be Part of a Common Response to Cell-Penetrating Peptides.Biomolecules. 2020 Nov 17;10(11):1567. doi: 10.3390/biom10111567. Biomolecules. 2020. PMID: 33213097 Free PMC article.
-
Reduced ribosomal DNA transcription in the prefrontal cortex of suicide victims: consistence of new molecular RT-qPCR findings with previous morphometric data from AgNOR-stained pyramidal neurons.Eur Arch Psychiatry Clin Neurosci. 2021 Apr;271(3):567-576. doi: 10.1007/s00406-021-01232-4. Epub 2021 Jan 26. Eur Arch Psychiatry Clin Neurosci. 2021. PMID: 33501518 Free PMC article.
References
-
- Parrish J. Z., Emoto K., Kim M. D., Jan Y. N. (2007) Annu. Rev. Neurosci. 30, 399–423 - PubMed
-
- Miller F. D., Kaplan D. R. (2003) Curr. Opin. Neurobiol. 13, 391–398 - PubMed
-
- Dijkhuizen P. A., Ghosh A. (2005) Prog. Brain Res. 147, 17–27 - PubMed
-
- Redmond L., Kashani A. H., Ghosh A. (2002) Neuron 34, 999–1010 - PubMed
-
- Wayman G. A., Impey S., Marks D., Saneyoshi T., Grant W. F., Derkach V., Soderling T. R. (2006) Neuron 50, 897–909 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

