RsgA releases RbfA from 30S ribosome during a late stage of ribosome biosynthesis
- PMID: 21102555
- PMCID: PMC3020115
- DOI: 10.1038/emboj.2010.291
RsgA releases RbfA from 30S ribosome during a late stage of ribosome biosynthesis
Abstract
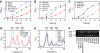
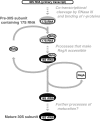
RsgA is a 30S ribosomal subunit-binding GTPase with an unknown function, shortage of which impairs maturation of the 30S subunit. We identified multiple gain-of-function mutants of Escherichia coli rbfA, the gene for a ribosome-binding factor, that suppress defects in growth and maturation of the 30S subunit of an rsgA-null strain. These mutations promote spontaneous release of RbfA from the 30S subunit, indicating that cellular disorders upon depletion of RsgA are due to prolonged retention of RbfA on the 30S subunit. We also found that RsgA enhances release of RbfA from the mature 30S subunit in a GTP-dependent manner but not from a precursor form of the 30S subunit. These findings indicate that the function of RsgA is to release RbfA from the 30S subunit during a late stage of ribosome biosynthesis. This is the first example of the action of a GTPase on the bacterial ribosome assembly described at the molecular level.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures




Similar articles
-
The C-terminal helix in the YjeQ zinc-finger domain catalyzes the release of RbfA during 30S ribosome subunit assembly.RNA. 2015 Jun;21(6):1203-16. doi: 10.1261/rna.049171.114. Epub 2015 Apr 22. RNA. 2015. PMID: 25904134 Free PMC article.
-
Lamotrigine-mediated rescue of RsgA-deficient Escherichia coli reveals another role of IF2 in ribosome biogenesis.J Bacteriol. 2024 Jul 25;206(7):e0011924. doi: 10.1128/jb.00119-24. Epub 2024 Jun 5. J Bacteriol. 2024. PMID: 38837341 Free PMC article.
-
Binding properties of YjeQ (RsgA), RbfA, RimM and Era to assembly intermediates of the 30S subunit.Nucleic Acids Res. 2016 Nov 16;44(20):9918-9932. doi: 10.1093/nar/gkw613. Epub 2016 Jul 5. Nucleic Acids Res. 2016. PMID: 27382067 Free PMC article.
-
Assembly of bacterial ribosomes.Annu Rev Biochem. 2011;80:501-26. doi: 10.1146/annurev-biochem-062608-160432. Annu Rev Biochem. 2011. PMID: 21529161 Review.
-
[Ribosome: lessons of a molecular factory construction].Mol Biol (Mosk). 2014 Jul-Aug;48(4):543-60. Mol Biol (Mosk). 2014. PMID: 25842841 Review. Russian.
Cited by
-
A conserved rRNA switch is central to decoding site maturation on the small ribosomal subunit.Sci Adv. 2021 Jun 4;7(23):eabf7547. doi: 10.1126/sciadv.abf7547. Print 2021 Jun. Sci Adv. 2021. PMID: 34088665 Free PMC article.
-
The cryo-EM structure of YjeQ bound to the 30S subunit suggests a fidelity checkpoint function for this protein in ribosome assembly.Proc Natl Acad Sci U S A. 2017 Apr 25;114(17):E3396-E3403. doi: 10.1073/pnas.1618016114. Epub 2017 Apr 10. Proc Natl Acad Sci U S A. 2017. PMID: 28396444 Free PMC article.
-
Ribosome assembly defects subvert initiation Factor3 mediated scrutiny of bona fide start signal.Nucleic Acids Res. 2019 Dec 2;47(21):11368-11386. doi: 10.1093/nar/gkz825. Nucleic Acids Res. 2019. PMID: 31586395 Free PMC article.
-
Molecular Basis of the Slow Growth of Mycoplasma hominis on Different Energy Sources.Front Cell Infect Microbiol. 2022 Jul 7;12:918557. doi: 10.3389/fcimb.2022.918557. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 35873139 Free PMC article.
-
Structural insights into the assembly of the 30S ribosomal subunit in vivo: functional role of S5 and location of the 17S rRNA precursor sequence.Protein Cell. 2014 May;5(5):394-407. doi: 10.1007/s13238-014-0044-1. Epub 2014 Mar 28. Protein Cell. 2014. PMID: 24671761 Free PMC article.
References
-
- Arigoni F, Talabot F, Peitsch M, Edgerton MD, Meldrum E, Allet E, Fish R, Jamotte T, Curchod ML, Loferer H (1998) A genome-based approach for the identification of essential bacterial genes. Nat Biotechnol 16: 851–856 - PubMed
-
- Britton RA (2009) Role of GTPases in bacterial ribosome assembly. Annu Rev Microbiol 63: 155–176 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

