Structural features of cytochromes P450 and ligands that affect drug metabolism as revealed by X-ray crystallography and NMR
- PMID: 21103389
- PMCID: PMC2987628
- DOI: 10.4155/fmc.10.229
Structural features of cytochromes P450 and ligands that affect drug metabolism as revealed by X-ray crystallography and NMR
Erratum in
- Future Med Chem. 2010 Oct;2(10):1612
Abstract
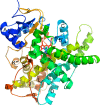
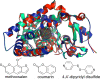
Cytochromes P450 (P450s) play a major role in the clearance of drugs, toxins, and environmental pollutants. Additionally, metabolism by P450s can result in toxic or carcinogenic products. The metabolism of pharmaceuticals by P450s is a major concern during the design of new drug candidates. Determining the interactions between P450s and compounds of very diverse structures is complicated by the variability in P450-ligand interactions. Understanding the protein structural elements and the chemical attributes of ligands that dictate their orientation in the P450 active site will aid in the development of effective and safe therapeutic agents. The goal of this review is to describe P450-ligand interactions from two perspectives. The first is the various structural elements that microsomal P450s have at their disposal to assume the different conformations observed in X-ray crystal structures. The second is P450-ligand dynamics analyzed by NMR relaxation studies.
Keywords: Cytochrome P450; NMR; P450; X-ray crystallography; drug metabolism; mammalian; microsomal; plasticity; protein structure; xenobiotic.
Figures




Similar articles
-
Structure and function of cytochromes P450 2B: from mechanism-based inactivators to X-ray crystal structures and back.Drug Metab Dispos. 2011 Jul;39(7):1113-21. doi: 10.1124/dmd.111.039719. Epub 2011 Apr 18. Drug Metab Dispos. 2011. PMID: 21502194 Free PMC article.
-
Analysis of cytochrome P450 CYP119 ligand-dependent conformational dynamics by two-dimensional NMR and X-ray crystallography.J Biol Chem. 2015 Apr 17;290(16):10000-17. doi: 10.1074/jbc.M114.627935. Epub 2015 Feb 10. J Biol Chem. 2015. PMID: 25670859 Free PMC article.
-
Crystal structures of human cytochrome P450 3A4 bound to metyrapone and progesterone.Science. 2004 Jul 30;305(5684):683-6. doi: 10.1126/science.1099736. Epub 2004 Jul 15. Science. 2004. PMID: 15256616
-
Conformational diversity and ligand tunnels of mammalian cytochrome P450s.Biotechnol Appl Biochem. 2013 Jan-Feb;60(1):134-45. doi: 10.1002/bab.1074. Biotechnol Appl Biochem. 2013. PMID: 23587001 Review.
-
Human P450s involved in drug metabolism and the use of structural modelling for understanding substrate selectivity and binding affinity.Xenobiotica. 2009 Aug;39(8):625-35. doi: 10.1080/00498250903000255. Xenobiotica. 2009. PMID: 19514836 Review.
Cited by
-
A large-scale allosteric transition in cytochrome P450 3A4 revealed by luminescence resonance energy transfer (LRET).PLoS One. 2013 Dec 23;8(12):e83898. doi: 10.1371/journal.pone.0083898. eCollection 2013. PLoS One. 2013. PMID: 24376769 Free PMC article.
-
β-Naphtoflavone and Ethanol Induce Cytochrome P450 and Protect towards MPP⁺ Toxicity in Human Neuroblastoma SH-SY5Y Cells.Int J Mol Sci. 2018 Oct 28;19(11):3369. doi: 10.3390/ijms19113369. Int J Mol Sci. 2018. PMID: 30373287 Free PMC article.
-
Methadone pharmacogenetics in vitro and in vivo: Metabolism by CYP2B6 polymorphic variants and genetic variability in paediatric disposition.Br J Clin Pharmacol. 2022 Nov;88(11):4881-4893. doi: 10.1111/bcp.15393. Epub 2022 Jun 20. Br J Clin Pharmacol. 2022. PMID: 35538637 Free PMC article.
-
Functional characterization of cytochromes P450 2B from the desert woodrat Neotoma lepida.Toxicol Appl Pharmacol. 2014 Feb 1;274(3):393-401. doi: 10.1016/j.taap.2013.12.005. Epub 2013 Dec 19. Toxicol Appl Pharmacol. 2014. PMID: 24361551 Free PMC article.
-
Assessment of glibenclamide pharmacokinetics in poloxamer 407-induced hyperlipidemic rats.Saudi Pharm J. 2021 Jul;29(7):719-723. doi: 10.1016/j.jsps.2021.05.002. Epub 2021 May 20. Saudi Pharm J. 2021. PMID: 34400867 Free PMC article.
References
-
- Kramer MA, Tracy TS. Studying cytochrome P450 kinetics in drug metabolism. Expert Opin Drug Metab Toxicol. 2008;4(5):591–603. - PubMed
-
- Domanski TL, Halpert JR. Analysis of mammalian cytochrome P450 structure and function by site-directed mutagenesis. Curr Drug Metab. 2001;2(2):117–137. - PubMed
-
- Guengerich FP, Parikh A, Yun C-H, et al. What makes P450s work? Searches for answers with known and new P450s. Drug Metab Rev. 2000;32(3–4):267–281. - PubMed
-
- de Graaf C, Vermeulen NP, Feenstra KA. Cytochrome P450 in silico: an integrative modeling approach. J Med Chem. 2005;48(8):2725–2755. - PubMed
-
- Lewis DF. Human P450s involved in drug metabolism and the use of structural modelling for understanding substrate selectivity and binding affinity. Xenobiotica. 2009;39(8):625–635. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
