Comparative transcriptome profiling analyses during the lag phase uncover YAP1, PDR1, PDR3, RPN4, and HSF1 as key regulatory genes in genomic adaptation to the lignocellulose derived inhibitor HMF for Saccharomyces cerevisiae
- PMID: 21106074
- PMCID: PMC3091778
- DOI: 10.1186/1471-2164-11-660
Comparative transcriptome profiling analyses during the lag phase uncover YAP1, PDR1, PDR3, RPN4, and HSF1 as key regulatory genes in genomic adaptation to the lignocellulose derived inhibitor HMF for Saccharomyces cerevisiae
Abstract
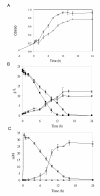
Background: The yeast Saccharomyces cerevisiae is able to adapt and in situ detoxify lignocellulose derived inhibitors such as furfural and HMF. The length of lag phase for cell growth in response to the inhibitor challenge has been used to measure tolerance of strain performance. Mechanisms of yeast tolerance at the genome level remain unknown. Using systems biology approach, this study investigated comparative transcriptome profiling, metabolic profiling, cell growth response, and gene regulatory interactions of yeast strains and selective gene deletion mutations in response to HMF challenges during the lag phase of growth.
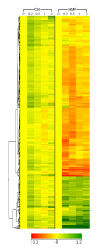
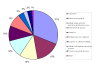
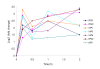
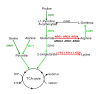
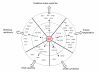
Results: We identified 365 candidate genes and found at least 3 significant components involving some of these genes that enable yeast adaptation and tolerance to HMF in yeast. First, functional enzyme coding genes such as ARI1, ADH6, ADH7, and OYE3, as well as gene interactions involved in the biotransformation and inhibitor detoxification were the direct driving force to reduce HMF damages in cells. Expressions of these genes were regulated by YAP1 and its closely related regulons. Second, a large number of PDR genes, mainly regulated by PDR1 and PDR3, were induced during the lag phase and the PDR gene family-centered functions, including specific and multiple functions involving cellular transport such as TPO1, TPO4, RSB1, PDR5, PDR15, YOR1, and SNQ2, promoted cellular adaptation and survival in order to cope with the inhibitor stress. Third, expressed genes involving degradation of damaged proteins and protein modifications such as SHP1 and SSA4, regulated by RPN4, HSF1, and other co-regulators, were necessary for yeast cells to survive and adapt the HMF stress. A deletion mutation strain Δrpn4 was unable to recover the growth in the presence of HMF.
Conclusions: Complex gene interactions and regulatory networks as well as co-regulations exist in yeast adaptation and tolerance to the lignocellulose derived inhibitor HMF. Both induced and repressed genes involving diversified functional categories are accountable for adaptation and energy rebalancing in yeast to survive and adapt the HMF stress during the lag phase of growth. Transcription factor genes YAP1, PDR1, PDR3, RPN4, and HSF1 appeared to play key regulatory rules for global adaptation in the yeast S. cerevisiae.
Figures
References
-
- Bothast RJ, Saha BC. Ethanol production from agricultural biomass substrate. Adv Appl Microbiol. 1997;44:261–286. full_text.
-
- Outlaw J, Collins K, Duffield J. Agriculture as a producer and consumer of energy. CAB International, Wallingford, UK; 2005.
-
- Liu ZL, Saha BC, Slininger PJ. In: Bioenergy. 1. Wall J, Harwood C, Demain A, editor. Washington, DC: ASM Press; 2008. Lignocellulose biomass conversion to ethanol by Saccharomyces; pp. 17–36.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

