Alcohol induces RNA polymerase III-dependent transcription through c-Jun by co-regulating TATA-binding protein (TBP) and Brf1 expression
- PMID: 21106530
- PMCID: PMC3024733
- DOI: 10.1074/jbc.M110.192955
Alcohol induces RNA polymerase III-dependent transcription through c-Jun by co-regulating TATA-binding protein (TBP) and Brf1 expression
Abstract
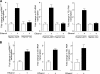
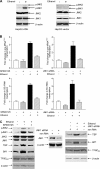
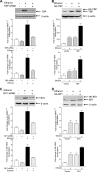
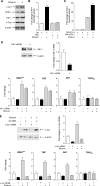
Chronic alcohol consumption is associated with steatohepatitis and cirrhosis, enhancing the risk for hepatocellular carcinoma. RNA polymerase (pol) III transcribes a variety of small, untranslated RNAs, including tRNAs and 5S rRNAs, which determine the biosynthetic capacity of cells. Increased RNA pol III-dependent transcription, observed in transformed cells and human tumors, is required for oncogenic transformation. Given that alcohol consumption increases risk for liver cancer, we examined whether alcohol regulates this class of genes. Ethanol induces RNA pol III-dependent transcription in both HepG2 cells and primary mouse hepatocytes in a manner that requires ethanol metabolism and the activation of JNK1. This regulatory event is mediated, at least in part, through the ability of ethanol to induce expression of the TFIIIB components, Brf1, and the TATA-binding protein (TBP). Induction of TBP, Brf1, and RNA pol III-dependent gene expression is driven by enhanced c-Jun expression. Ethanol promotes a marked increase in the direct recruitment of c-Jun to TBP, Brf1, and tRNA gene promoters. Chronic alcohol administration in mice leads to enhanced expression of TBP, Brf1, tRNA, and 5S rRNA gene transcription in the liver. These alcohol-dependent increases are more pronounced in transgenic animals that express the HCV NS5A protein that display increased incidence of liver tumors. Together, these results identify a new class of genes that are regulated by alcohol through the co-regulation of TFIIIB components and define a central role for c-Jun in this process.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

