A novel non-SET domain multi-subunit methyltransferase required for sequential nucleosomal histone H3 methylation by the mixed lineage leukemia protein-1 (MLL1) core complex
- PMID: 21106533
- PMCID: PMC3030342
- DOI: 10.1074/jbc.M110.174524
A novel non-SET domain multi-subunit methyltransferase required for sequential nucleosomal histone H3 methylation by the mixed lineage leukemia protein-1 (MLL1) core complex
Erratum in
- J Biol Chem. 2011 May 20;286(20):18344
Abstract
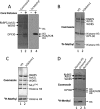
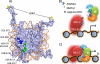
Gene expression within the context of eukaryotic chromatin is regulated by enzymes that catalyze histone lysine methylation. Histone lysine methyltransferases that have been identified to date possess the evolutionarily conserved SET or Dot1-like domains. We previously reported the identification of a new multi-subunit histone H3 lysine 4 methyltransferase lacking homology to the SET or Dot1 family of histone lysine methyltransferases. This enzymatic activity requires a complex that includes WRAD (WDR5, RbBP5, Ash2L, and DPY-30), a complex that is part of the MLL1 (mixed lineage leukemia protein-1) core complex but that also exists independently of MLL1 in the cell. Here, we report that the minimal complex required for WRAD enzymatic activity includes WDR5, RbBP5, and Ash2L and that DPY-30, although not required for enzymatic activity, increases the histone substrate specificity of the WRAD complex. We also show that WRAD requires zinc for catalytic activity, displays Michaelis-Menten kinetics, and is inhibited by S-adenosyl-homocysteine. In addition, we demonstrate that WRAD preferentially methylates lysine 4 of histone H3 within the context of the H3/H4 tetramer but does not methylate nucleosomal histone H3 on its own. In contrast, we find that MLL1 and WRAD are required for nucleosomal histone H3 methylation, and we provide evidence suggesting that each plays distinct structural and catalytic roles in the recognition and methylation of a nucleosome substrate. Our results indicate that WRAD is a new H3K4 methyltransferase with functions that include regulating the substrate and product specificities of the MLL1 core complex.
Figures


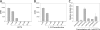


References
-
- Santos-Rosa H., Schneider R., Bernstein B. E., Karabetsou N., Morillon A., Weise C., Schreiber S. L., Mellor J., Kouzarides T. (2003) Mol. Cell 12, 1325–1332 - PubMed
-
- Cosgrove M. S., Wolberger C. (2005) Biochem. Cell Biol. 83, 468–476 - PubMed
-
- Wysocka J., Swigut T., Xiao H., Milne T. A., Kwon S. Y., Landry J., Kauer M., Tackett A. J., Chait B. T., Badenhorst P., Wu C., Allis C. D. (2006) Nature 442, 86–90 - PubMed
-
- Santos-Rosa H., Schneider R., Bannister A. J., Sherriff J., Bernstein B. E., Emre N. C., Schreiber S. L., Mellor J., Kouzarides T. (2002) Nature 419, 407–411 - PubMed
-
- Ruthenburg A. J., Allis C. D., Wysocka J. (2007) Mol. Cell 25, 15–30 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

