Assessing the potential for AAV vector genotoxicity in a murine model
- PMID: 21106988
- PMCID: PMC3069673
- DOI: 10.1182/blood-2010-08-302729
Assessing the potential for AAV vector genotoxicity in a murine model
Erratum in
- Blood. 2011 Jun 16;117(24):6739
Abstract
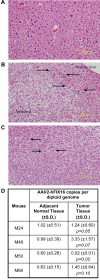
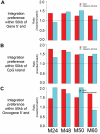
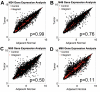
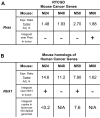
Gene transfer using adeno-associated virus (AAV) vectors has great potential for treating human disease. Recently, questions have arisen about the safety of AAV vectors, specifically, whether integration of vector DNA in transduced cell genomes promotes tumor formation. This study addresses these questions with high-dose liver-directed AAV-mediated gene transfer in the adult mouse as a model (80 AAV-injected mice and 52 controls). After 18 months of follow-up, AAV-injected mice did not show a significantly higher rate of hepatocellular carcinoma compared with controls. Tumors in mice treated with AAV vectors did not have significantly different amounts of vector DNA compared with adjacent normal tissue. A novel high-throughput method for identifying AAV vector integration sites was developed and used to clone 1029 integrants. Integration patterns in tumor tissue and adjacent normal tissue were similar to each other, showing preferences for active genes, cytosine-phosphate-guanosine islands, and guanosine/cytosine-rich regions. [corrected] Gene expression data showed that genes near integration sites did not show significant changes in expression patterns compared with genes more distal to integration sites. No integration events were identified as causing increased oncogene expression. Thus, we did not find evidence that AAV vectors cause insertional activation of oncogenes and subsequent tumor formation.
Figures

Comment in
-
Quest for safety at AAValon.Blood. 2011 Mar 24;117(12):3249-50. doi: 10.1182/blood-2011-02-321984. Blood. 2011. PMID: 21436076 No abstract available.
References
-
- Manno CS, Pierce GF, Arruda VR, et al. Successful transduction of liver in hemophilia by AAV-factor IX and limitations imposed by the host immune response. Nat Med. 2006;12(5):342–347. - PubMed
-
- Miller DG, Petek LM, Russell DW. Adeno-associated virus vectors integrate at chromosome breakage sites. Nat Genet. 2004;36(7):767–773. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

