Reverse type I inhibitor of Mycobacterium tuberculosis CYP125A1
- PMID: 21109436
- PMCID: PMC3011832
- DOI: 10.1016/j.bmcl.2010.11.007
Reverse type I inhibitor of Mycobacterium tuberculosis CYP125A1
Abstract
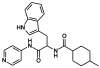
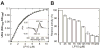
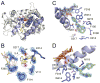
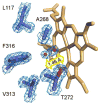
Cytochrome P450 CYP125A1 of Mycobacterium tuberculosis, a potential therapeutic target for tuberculosis in humans, initiates degradation of the aliphatic chain of host cholesterol and is essential for establishing M. tuberculosis infection in a mouse model of disease. We explored the interactions of CYP125A1 with a reverse type I inhibitor by X-ray structure analysis and UV-vis spectroscopy. Compound LP10 (α-[(4-methylcyclohexyl)carbonyl amino]-N-4-pyridinyl-1H-indole-3-propanamide), previously identified as a potent type II inhibitor of Trypanosomacruzi CYP51, shifts CYP125A1 to a water-coordinated low-spin state upon binding with low micromolar affinity. When LP10 is present in the active site, the crystal structure and spectral characteristics both demonstrate changes in lipophilic and electronic properties favoring coordination of the iron axial water ligand. These results provide an insight into the structural requirements for developing selective CYP125A1 inhibitors.
Copyright © 2010 Elsevier Ltd. All rights reserved.
Figures
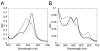
Similar articles
-
Potential drug targets in the Mycobacterium tuberculosis cytochrome P450 system.J Inorg Biochem. 2018 Mar;180:235-245. doi: 10.1016/j.jinorgbio.2018.01.010. Epub 2018 Jan 12. J Inorg Biochem. 2018. PMID: 29352597 Free PMC article. Review.
-
A comparison of steroid and lipid binding cytochrome P450s from Mycobacterium marinum and Mycobacterium tuberculosis.J Inorg Biochem. 2020 Aug;209:111116. doi: 10.1016/j.jinorgbio.2020.111116. Epub 2020 May 20. J Inorg Biochem. 2020. PMID: 32473484
-
Small-molecule scaffolds for CYP51 inhibitors identified by high-throughput screening and defined by X-ray crystallography.Antimicrob Agents Chemother. 2007 Nov;51(11):3915-23. doi: 10.1128/AAC.00311-07. Epub 2007 Sep 10. Antimicrob Agents Chemother. 2007. PMID: 17846131 Free PMC article.
-
Mycobacterium tuberculosis CYP130: crystal structure, biophysical characterization, and interactions with antifungal azole drugs.J Biol Chem. 2008 Feb 22;283(8):5069-80. doi: 10.1074/jbc.M708734200. Epub 2007 Dec 18. J Biol Chem. 2008. PMID: 18089574 Free PMC article.
-
Structural biology and biochemistry of cytochrome P450 systems in Mycobacterium tuberculosis.Drug Metab Rev. 2008;40(3):427-46. doi: 10.1080/03602530802186389. Drug Metab Rev. 2008. PMID: 18642141 Review.
Cited by
-
Drug modulation of water-heme interactions in low-spin P450 complexes of CYP2C9d and CYP125A1.Biochemistry. 2015 Feb 10;54(5):1198-207. doi: 10.1021/bi501402k. Epub 2015 Jan 29. Biochemistry. 2015. PMID: 25591012 Free PMC article.
-
Potential drug targets in the Mycobacterium tuberculosis cytochrome P450 system.J Inorg Biochem. 2018 Mar;180:235-245. doi: 10.1016/j.jinorgbio.2018.01.010. Epub 2018 Jan 12. J Inorg Biochem. 2018. PMID: 29352597 Free PMC article. Review.
-
Cholesterol metabolism: a potential therapeutic target in Mycobacteria.Br J Pharmacol. 2017 Jul;174(14):2194-2208. doi: 10.1111/bph.13694. Epub 2017 Jan 24. Br J Pharmacol. 2017. PMID: 28002883 Free PMC article. Review.
-
Fragment-Based Approaches to the Development of Mycobacterium tuberculosis CYP121 Inhibitors.J Med Chem. 2016 Apr 14;59(7):3272-302. doi: 10.1021/acs.jmedchem.6b00007. Epub 2016 Mar 22. J Med Chem. 2016. PMID: 27002486 Free PMC article.
-
Roles of cysteine in the structure and metabolic function of Mycobacterium tuberculosis CYP142A1.RSC Adv. 2022 Aug 30;12(38):24447-24455. doi: 10.1039/d2ra04257f. eCollection 2022 Aug 30. RSC Adv. 2022. PMID: 36128375 Free PMC article.
References
-
- Belin P, Le Du MH, Fielding A, Lequin O, Jacquet M, Charbonnier J-B, Lecoq A, Thai R, Courcon M, Masson C, Dugave C, Genet R, Pernodet J-L, Gondry M. Identification and structural basis of the reaction catalyzed by CYP121, an essential cytochrome P450 in Mycobacterium tuberculosis. Proc Natl Acad Sci U S A. 2009;106:7426–7431. - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Chemical Information

