Dynamics in the mixed microbial concourse
- PMID: 21123647
- PMCID: PMC2994034
- DOI: 10.1101/gad.1985210
Dynamics in the mixed microbial concourse
Abstract
Isolated, clonal populations of cells are rarely found in nature. The emergent properties of microbial consortia present a challenge for the systems approach to biology, as chances for competition, communication, or collaboration multiply with the number of interacting agents. This review focuses on recent work on intercourse within biofilms, among quorum-sensing populations, and between cross-feeding metabolic cooperators. New tools from synthetic biology allow microbial interactions to be designed and tightly controlled, creating valuable model systems. We address both natural and synthetic partnerships, with an emphasis on how system behaviors derive from the properties of their components. Essential features of microbial biology arose in the context of a very mixed culture and cannot be understood without unscrambling it.
Figures


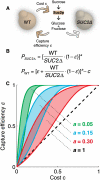
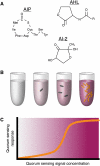

References
-
- Alon U 2007. An introduction to systems biology: Design principles of biological circuits. CRC Press, Boca Raton, FL
-
- Atkinson S, Chang CY, Patrick HL, Buckley CM, Wang Y, Sockett RE, Cámara M, Williams P 2008. Functional interplay between the Yersinia pseudotuberculosis YpsRI and YtbRI quorum sensing systems modulates swimming motility by controlling expression of flhDC and fliA. Mol Microbiol 69: 137–151 - PubMed
-
- Balagaddé FK, You L, Hansen CL, Arnold FH, Quake SR 2005. Long-term monitoring of bacteria undergoing programmed population control in a microchemostat. Science 309: 137–140 - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
