Abundances and potential activities of nitrogen cycling microbial communities along a chronosequence of a glacier forefield
- PMID: 21124490
- PMCID: PMC3131848
- DOI: 10.1038/ismej.2010.184
Abundances and potential activities of nitrogen cycling microbial communities along a chronosequence of a glacier forefield
Abstract
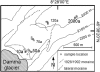
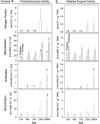
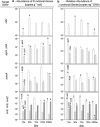
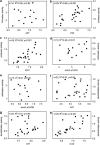
Glacier forefields are ideal ecosystems to study the development of nutrient cycles as well as single turnover processes during soil development. In this study, we examined the ecology of the microbial nitrogen (N) cycle in bulk soil samples from a chronosequence of the Damma glacier, Switzerland. Major processes of the N cycle were reconstructed on the genetic as well as the potential enzyme activity level at sites of the chronosequence that have been ice-free for 10, 50, 70, 120 and 2000 years. In our study, we focused on N fixation, mineralization (chitinolysis and proteolysis), nitrification and denitrification. Our results suggest that mineralization, mainly the decomposition of deposited organic material, was the main driver for N turnover in initial soils, that is, ice-free for 10 years. Transient soils being ice-free for 50 and 70 years were characterized by a high abundance of N fixing microorganisms. In developed soils, ice-free for 120 and 2000 years, significant rates of nitrification and denitrification were measured. Surprisingly, copy numbers of the respective functional genes encoding the corresponding enzymes were already high in the initial phase of soil development. This clearly indicates that the genetic potential is not the driver for certain functional traits in the initial phase of soil formation but rather a well-balanced expression of the respective genes coding for selected functions.
Figures
Similar articles
-
The Spatial Dynamics of Diazotrophs in the Forefield of Three Tibetan Glaciers.Microb Ecol. 2024 Nov 13;87(1):138. doi: 10.1007/s00248-024-02460-6. Microb Ecol. 2024. PMID: 39532738 Free PMC article.
-
Abundance of microbes involved in nitrogen transformation in the rhizosphere of Leucanthemopsis alpina (L.) Heywood grown in soils from different sites of the Damma glacier forefield.Microb Ecol. 2010 Nov;60(4):762-70. doi: 10.1007/s00248-010-9695-5. Epub 2010 Jun 12. Microb Ecol. 2010. PMID: 20549199
-
Changes in diversity and functional gene abundances of microbial communities involved in nitrogen fixation, nitrification, and denitrification in a tidal wetland versus paddy soils cultivated for different time periods.Appl Environ Microbiol. 2011 Sep;77(17):6109-16. doi: 10.1128/AEM.01751-10. Epub 2011 Jul 15. Appl Environ Microbiol. 2011. PMID: 21764972 Free PMC article.
-
Nitrogen turnover in soil and global change.FEMS Microbiol Ecol. 2011 Oct;78(1):3-16. doi: 10.1111/j.1574-6941.2011.01165.x. Epub 2011 Jul 29. FEMS Microbiol Ecol. 2011. PMID: 21707675 Review.
-
Microbial Biogeochemical Cycling of Nitrogen in Arid Ecosystems.Microbiol Mol Biol Rev. 2022 Jun 15;86(2):e0010921. doi: 10.1128/mmbr.00109-21. Epub 2022 Apr 7. Microbiol Mol Biol Rev. 2022. PMID: 35389249 Free PMC article. Review.
Cited by
-
Variation in denitrifying bacterial communities along a primary succession in the Hailuogou Glacier retreat area, China.PeerJ. 2019 Aug 14;7:e7356. doi: 10.7717/peerj.7356. eCollection 2019. PeerJ. 2019. PMID: 31428538 Free PMC article.
-
Dynamics of biochemical properties associated with soil nitrogen mineralization following nitrification inhibitor and fungicide applications.Environ Sci Pollut Res Int. 2017 Apr;24(12):11340-11348. doi: 10.1007/s11356-017-8762-6. Epub 2017 Mar 17. Environ Sci Pollut Res Int. 2017. PMID: 28303541
-
Do Aerial Nitrogen Depositions Affect Fungal and Bacterial Communities of Oak Leaves?Front Microbiol. 2021 Apr 16;12:633535. doi: 10.3389/fmicb.2021.633535. eCollection 2021. Front Microbiol. 2021. PMID: 33935994 Free PMC article.
-
The Spatial Dynamics of Diazotrophs in the Forefield of Three Tibetan Glaciers.Microb Ecol. 2024 Nov 13;87(1):138. doi: 10.1007/s00248-024-02460-6. Microb Ecol. 2024. PMID: 39532738 Free PMC article.
-
The Root-Associated Microbial Community of the World's Highest Growing Vascular Plants.Microb Ecol. 2016 Aug;72(2):394-406. doi: 10.1007/s00248-016-0779-8. Epub 2016 May 31. Microb Ecol. 2016. PMID: 27245598 Free PMC article.
References
-
- Babic KH, Schauss K, Hai B, Sikora S, Redzepovic S, Radl V, et al. Influence of different Sinorhizobium meliloti inocula on abundance of genes involved in nitrogen transformations in the rhizosphere of alfalfa (Medicago sativa L.) Environ Microbiol. 2008;10:2922–2930. - PubMed
-
- Bach HJ, Hartmann A, Schloter M, Munch JC. PCR primers and functional probes for amplification and detection of bacterial genes for extracellular peptidases in single strains and in soil. J Microbiol Meth. 2001;44:173–182. - PubMed
-
- Bardgett RD, Walker LR. Impact of coloniser plant species on the development of decomposer microbial communities following deglaciation. Soil Biol Biochem. 2004;36:555–559.
-
- Bernasconi SM. Weathering, soil formation and initial ecosystem evolution on a glacier forefield: a case study from the Damma Glacier, Switzerland. Mineral Mag. 2008;72:19–22.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

