The parental non-equivalence of imprinting control regions during mammalian development and evolution
- PMID: 21124941
- PMCID: PMC2987832
- DOI: 10.1371/journal.pgen.1001214
The parental non-equivalence of imprinting control regions during mammalian development and evolution
Abstract
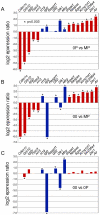
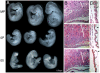
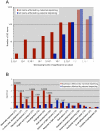
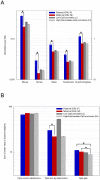
In mammals, imprinted gene expression results from the sex-specific methylation of imprinted control regions (ICRs) in the parental germlines. Imprinting is linked to therian reproduction, that is, the placenta and imprinting emerged at roughly the same time and potentially co-evolved. We assessed the transcriptome-wide and ontology effect of maternally versus paternally methylated ICRs at the developmental stage of setting of the chorioallantoic placenta in the mouse (8.5dpc), using two models of imprinting deficiency including completely imprint-free embryos. Paternal and maternal imprints have a similar quantitative impact on the embryonic transcriptome. However, transcriptional effects of maternal ICRs are qualitatively focused on the fetal-maternal interface, while paternal ICRs weakly affect non-convergent biological processes, with little consequence for viability at 8.5dpc. Moreover, genes regulated by maternal ICRs indirectly influence genes regulated by paternal ICRs, while the reverse is not observed. The functional dominance of maternal imprints over early embryonic development is potentially linked to selection pressures favoring methylation-dependent control of maternal over paternal ICRs. We previously hypothesized that the different methylation histories of ICRs in the maternal versus the paternal germlines may have put paternal ICRs under higher mutational pressure to lose CpGs by deamination. Using comparative genomics of 17 extant mammalian species, we show here that, while ICRs in general have been constrained to maintain more CpGs than non-imprinted sequences, the rate of CpG loss at paternal ICRs has indeed been higher than at maternal ICRs during evolution. In fact, maternal ICRs, which have the characteristics of CpG-rich promoters, have gained CpGs compared to non-imprinted CpG-rich promoters. Thus, the numerical and, during early embryonic development, functional dominance of maternal ICRs can be explained as the consequence of two orthogonal evolutionary forces: pressure to tightly regulate genes affecting the fetal-maternal interface and pressure to avoid the mutagenic environment of the paternal germline.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Comment in
-
Methylation from mother.Nat Rev Genet. 2011 Jan;12(1):6. doi: 10.1038/nrg2923. Epub 2010 Dec 7. Nat Rev Genet. 2011. PMID: 21135864 No abstract available.
References
-
- Wood AJ, Oakey RJ. Genomic imprinting in mammals: emerging themes and established theories. PLoS Genet. 2006;2:e147. doi: 10.1371/journal.pgen.0020147. - DOI - PMC - PubMed
-
- Tucker KL, Beard C, Dausmann J, Jackson-Grusby L, Laird PW, et al. Germ-line passage is required for establishment of methylation and expression patterns of imprinted but not of nonimprinted genes. Genes Dev. 1996;10:1008–1020. - PubMed
-
- Bourc'his D, Xu GL, Lin CS, Bollman B, Bestor TH. Dnmt3L and the establishment of maternal genomic imprints. Science. 2001;294:2536–2539. - PubMed
-
- Kaneda M, Okano M, Hata K, Sado T, Tsujimoto N, et al. Essential role for de novo DNA methyltransferase Dnmt3a in paternal and maternal imprinting. Nature. 2004;429:900–903. - PubMed
-
- Bourc'his D, Bestor TH. Origins of extreme sexual dimorphism in genomic imprinting. Cytogenet Genome Res. 2006;113:36–40. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

