miRNAs in human cancer
- PMID: 21125669
- PMCID: PMC3069496
- DOI: 10.1002/path.2806
miRNAs in human cancer
Abstract
Mature microRNAs (miRNAs) are single-stranded RNA molecules of 20-23 nucleotide (nt) length that control gene expression in many cellular processes. These molecules typically reduce the stability of mRNAs, including those of genes that mediate processes in tumorigenesis, such as inflammation, cell cycle regulation, stress response, differentiation, apoptosis and invasion. miRNA targeting is mostly achieved through specific base-pairing interactions between the 5' end ('seed' region) of the miRNA and sites within coding and untranslated regions (UTRs) of mRNAs; target sites in the 3' UTR lead to more effective mRNA destabilization. Since miRNAs frequently target hundreds of mRNAs, miRNA regulatory pathways are complex. To provide a critical overview of miRNA dysregulation in cancer, we first discuss the methods currently available for studying the role of miRNAs in cancer and then review miRNA genomic organization, biogenesis and mechanism of target recognition, examining how these processes are altered in tumorigenesis. Given the critical role miRNAs play in tumorigenesis processes and their disease-specific expression, they hold potential as therapeutic targets and novel biomarkers.
Copyright © 2010 Pathological Society of Great Britain and Ireland. Published by John Wiley & Sons, Ltd.
Figures
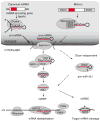


References
-
- Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75:843–854. - PubMed
-
- Reinhart BJ, Slack FJ, Basson M, et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature. 2000;403:901–906. - PubMed
-
- Wightman B, Ha I, Ruvkun G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell. 1993;75:855–862. - PubMed
-
- Wightman B, Burglin TR, Gatto J, et al. Negative regulatory sequences in the lin-14 3′-untranslated region are necessary to generate a temporal switch during Caenorhabditis elegans development. Genes Dev. 1991;5:1813–1824. - PubMed
-
- Lagos-Quintana M, Rauhut R, Lendeckel W, et al. Identification of Novel Genes Coding for Small Expressed RNAs. Science. 2001;294:853–858. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

