Positive selection at high temperature reduces gene transcription in the bacteriophage ϕX174
- PMID: 21129199
- PMCID: PMC3003669
- DOI: 10.1186/1471-2148-10-378
Positive selection at high temperature reduces gene transcription in the bacteriophage ϕX174
Abstract
Background: Gene regulation plays a central role in the adaptation of organisms to their environments. There are many molecular components to gene regulation, and it is often difficult to determine both the genetic basis of adaptation and the evolutionary forces that influence regulation. In multiple evolution experiments with the bacteriophage ϕX174, adaptive substitutions in cis-acting regulatory sequences sweep through the phage population as the result of strong positive selection at high temperatures that are non-permissive for laboratory-adapted phage. For one cis-regulatory region, we investigate the individual effects of four adaptive substitutions on transcript levels and fitness for phage growing on three hosts at two temperatures.
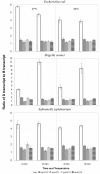
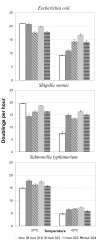
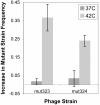
Results: The effect of the four individual substitutions on transcript levels is to down-regulate gene expression, regardless of temperature or host. To ascertain the conditions under which these substitutions are adaptive, fitness was measured by a variety of methods for several bacterial hosts growing at two temperatures, the control temperature of 37°C and the selective temperature of 42°C. Time to lysis and doublings per hour indicate that the four substitutions individually improve fitness over the ancestral strain at high temperature independent of the bacterial host in which the fitness was measured. Competition assays between the ancestral strain and either of two mutant strains indicate that both mutants out-compete the ancestor at high temperature, but the relative frequencies of each phage remain the same at the control temperature.
Conclusions: Our results strongly suggest that gene transcription plays an important role in influencing fitness in the bacteriophage ϕX174, and different point mutations in a single cis-regulatory region provided the genetic basis for this role in adaptation to high temperature. We speculate that the adaptive nature of these substitutions is due to the physiology of the host at high temperature or the need to maintain particular ratios of phage proteins during capsid assembly. Our investigation of regulatory evolution contributes to interpreting genome-level assessments of regulatory variation, as well as to understanding the molecular basis of adaptation.
Figures

References
-
- Li WH, Wu CI, Luo CC. A new method for estimating synonymous and nonsynonymous rates of nucleotide substitution considering the relative likelihood of nucleotide and codon changes. Mol Biol Evol. 1985;2(2):150–174. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

