Translation on demand by a simple RNA-based thermosensor
- PMID: 21131278
- PMCID: PMC3074152
- DOI: 10.1093/nar/gkq1252
Translation on demand by a simple RNA-based thermosensor
Abstract
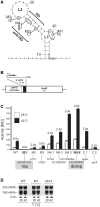
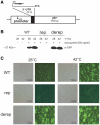
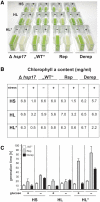
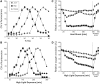
Structured RNA regions are important gene control elements in prokaryotes and eukaryotes. Here, we show that the mRNA of a cyanobacterial heat shock gene contains a built-in thermosensor critical for photosynthetic activity under stress conditions. The exceptionally short 5'-untranslated region is comprised of a single hairpin with an internal asymmetric loop. It inhibits translation of the Synechocystis hsp17 transcript at normal growth conditions, permits translation initiation under stress conditions and shuts down Hsp17 production in the recovery phase. Point mutations that stabilized or destabilized the RNA structure deregulated reporter gene expression in vivo and ribosome binding in vitro. Introduction of such point mutations into the Synechocystis genome produced severe phenotypic defects. Reversible formation of the open and closed structure was beneficial for viability, integrity of the photosystem and oxygen evolution. Continuous production of Hsp17 was detrimental when the stress declined indicating that shutting-off heat shock protein production is an important, previously unrecognized function of RNA thermometers. We discovered a simple biosensor that strictly adjusts the cellular level of a molecular chaperone to the physiological need.
Figures




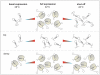
References
-
- Castielli O, De la Cerda B, Navarro JA, Hervas M, De la Rosa MA. Proteomic analyses of the response of cyanobacteria to different stress conditions. FEBS Lett. 2009;583:1753–1758. - PubMed
-
- Latifi A, Ruiz M, Zhang CC. Oxidative stress in cyanobacteria. FEMS Microbiol. Rev. 2009;33:258–278. - PubMed
-
- Lee S, Prochaska DJ, Fang F, Barnum SR. A 16.6-kilodalton protein in the Cyanobacterium Synechocystis sp. PCC 6803 plays a role in the heat shock response. Curr. Microbiol. 1998;37:403–407. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

