Candida albicans Hap43 is a repressor induced under low-iron conditions and is essential for iron-responsive transcriptional regulation and virulence
- PMID: 21131439
- PMCID: PMC3067405
- DOI: 10.1128/EC.00158-10
Candida albicans Hap43 is a repressor induced under low-iron conditions and is essential for iron-responsive transcriptional regulation and virulence
Abstract
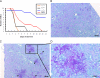
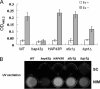
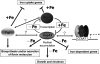
Candida albicans is an opportunistic fungal pathogen that exists as normal flora in healthy human bodies but causes life-threatening infections in immunocompromised patients. In addition to innate and adaptive immunities, hosts also resist microbial infections by developing a mechanism of "natural resistance" that maintains a low level of free iron to restrict the growth of invading pathogens. C. albicans must overcome this iron-deprived environment to cause infections. There are three types of iron-responsive transcriptional regulators in fungi; Aft1/Aft2 activators in yeast, GATA-type repressors in many fungi, and HapX/Php4 in Schizosaccharomyces pombe and Aspergillus species. In this study, we characterized the iron-responsive regulator Hap43, which is the C. albicans homolog of HapX/Php4 and is repressed by the GATA-type repressor Sfu1 under iron-sufficient conditions. We provide evidence that Hap43 is essential for the growth of C. albicans under low-iron conditions and for C. albicans virulence in a mouse model of infection. Hap43 was not required for iron acquisition under low-iron conditions. Instead, it was responsible for repression of genes that encode iron-dependent proteins involved in mitochondrial respiration and iron-sulfur cluster assembly. We also demonstrated that Hap43 executes its function by becoming a transcriptional repressor and accumulating in the nucleus in response to iron deprivation. Finally, we found a connection between Hap43 and the global corepressor Tup1 in low-iron-induced flavinogenesis. Taken together, our data suggest a complex interplay among Hap43, Sfu1, and Tup1 to coordinately regulate iron acquisition, iron utilization, and other iron-responsive metabolic activities.
Figures

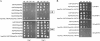




Similar articles
-
Diverse Hap43-independent functions of the Candida albicans CCAAT-binding complex.Eukaryot Cell. 2013 Jun;12(6):804-15. doi: 10.1128/EC.00014-13. Epub 2013 Mar 29. Eukaryot Cell. 2013. PMID: 23543673 Free PMC article.
-
An iron homeostasis regulatory circuit with reciprocal roles in Candida albicans commensalism and pathogenesis.Cell Host Microbe. 2011 Aug 18;10(2):118-35. doi: 10.1016/j.chom.2011.07.005. Cell Host Microbe. 2011. PMID: 21843869 Free PMC article.
-
Expression of Candida albicans Sfu1 in fission yeast complements the loss of the iron-regulatory transcription factor Fep1 and requires Tup co-repressors.Yeast. 2007 Oct;24(10):883-900. doi: 10.1002/yea.1539. Yeast. 2007. PMID: 17724773
-
Coordinated regulation of iron metabolism in Cryptococcus neoformans by GATA and CCAAT transcription factors: connections with virulence.Curr Genet. 2021 Aug;67(4):583-593. doi: 10.1007/s00294-021-01172-5. Epub 2021 Mar 24. Curr Genet. 2021. PMID: 33760942 Free PMC article. Review.
-
Mechanisms and regulation of iron uptake and the role of iron in pathogenesis of Candida albicans.Crit Rev Microbiol. 2025 May 24:1-18. doi: 10.1080/1040841X.2025.2510256. Online ahead of print. Crit Rev Microbiol. 2025. PMID: 40411301 Review.
Cited by
-
The adaptive response to iron involves changes in energetic strategies in the pathogen Candida albicans.Microbiologyopen. 2020 Feb;9(2):e970. doi: 10.1002/mbo3.970. Epub 2019 Dec 1. Microbiologyopen. 2020. PMID: 31788966 Free PMC article.
-
Experimental evolution reveals a general role for the methyltransferase Hmt1 in noise buffering.PLoS Biol. 2019 Oct 15;17(10):e3000433. doi: 10.1371/journal.pbio.3000433. eCollection 2019 Oct. PLoS Biol. 2019. PMID: 31613873 Free PMC article.
-
The Iron-Dependent Regulation of the Candida albicans Oxidative Stress Response by the CCAAT-Binding Factor.PLoS One. 2017 Jan 25;12(1):e0170649. doi: 10.1371/journal.pone.0170649. eCollection 2017. PLoS One. 2017. PMID: 28122000 Free PMC article.
-
The Fungal Pathogen Candida glabrata Does Not Depend on Surface Ferric Reductases for Iron Acquisition.Front Microbiol. 2017 Jun 8;8:1055. doi: 10.3389/fmicb.2017.01055. eCollection 2017. Front Microbiol. 2017. PMID: 28642757 Free PMC article.
-
Candida albicans Hap43 Domains Are Required under Iron Starvation but Not Excess.Front Microbiol. 2017 Dec 1;8:2388. doi: 10.3389/fmicb.2017.02388. eCollection 2017. Front Microbiol. 2017. PMID: 29250054 Free PMC article.
References
-
- Archibald F. 1983. Lactobacillus plantarum, an organism not requiring iron. FEMS Microbiol. Lett. 19:29–32
-
- Blaiseau P. L., Lesuisse E., Camadro J. M. 2001. Aft2p, a novel iron-regulated transcription activator that modulates, with Aft1p, intracellular iron use and resistance to oxidative stress in yeast. J. Biol. Chem. 276:34221–34226 - PubMed
-
- Boretsky Y. R., et al. 2005. Positive selection of mutants defective in transcriptional repression of riboflavin synthesis by iron in the flavinogenic yeast Pichia guilliermondii. FEMS Yeast Res. 5:829–837 - PubMed
-
- Bourgarel D., Nguyen C. C., Bolotin-Fukuhara M. 1999. HAP4, the glucose-repressed regulated subunit of the HAP transcriptional complex involved in the fermentation-respiration shift, has a functional homologue in the respiratory yeast Kluyveromyces lactis. Mol. Microbiol. 31:1205–1215 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

