Immunohistochemical methods for predicting cell of origin and survival in patients with diffuse large B-cell lymphoma treated with rituximab
- PMID: 21135273
- PMCID: PMC3058275
- DOI: 10.1200/JCO.2010.30.0368
Immunohistochemical methods for predicting cell of origin and survival in patients with diffuse large B-cell lymphoma treated with rituximab
Abstract
Purpose: Patients with diffuse large B-cell lymphoma (DLBCL) can be divided into prognostic groups based on the cell of origin of the tumor as determined by microarray analysis. Various immunohistochemical algorithms have been developed to replicate these microarray results and/or stratify patients according to survival. This study compares some of those algorithms and also proposes some modifications.
Patients and methods: Two-hundred and sixty-two cases of de novo DLBCL treated with rituximab and cyclophosphamide, doxorubicin, vincristine, and prednisone (CHOP) or CHOP-like therapy were examined.
Results: The Choi algorithm and Hans algorithm had high concordance with the microarray results. Modifications of the Choi and Hans algorithms for ease of use still retained high concordance with the microarray results. Although the Nyman and Muris algorithms had high concordance with the microarray results, each had a low value for either sensitivity or specificity. The use of LMO2 alone showed the lowest concordance with the microarray results. A new algorithm (Tally) using a combination of antibodies, but without regard to the order of examination, showed the greatest concordance with microarray results. All of the algorithms divided patients into groups with significantly different overall and event-free survivals, but with different hazard ratios. With the exception of the Nyman algorithm, this survival prediction was independent of the International Prognostic Index. Although the Muris algorithm had prognostic significance, it misclassified a large number of cases with activated B-cell type DLBCL.
Conclusion: The Tally algorithm showed the best concordance with the microarray data while maintaining prognostic significance and ease of use.
Conflict of interest statement
Authors' disclosures of potential conflicts of interest and author contributions are found at the end of this article.
Figures

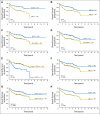
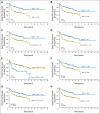

References
-
- Alizadeh AA, Eisen MB, Davis RE, et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature. 2000;403:503–511. - PubMed
-
- Hans CP, Weisenburger DD, Greiner TC, et al. Confirmation of the molecular classification of diffuse large B-cell lymphoma by immunohistochemistry using a tissue microarray. Blood. 2004;103:275–282. - PubMed
-
- Muris JJ, Meijer CJ, Vos W, et al. Immunohistochemical profiling based on Bcl-2, CD10 and MUM1 expression improves risk stratification in patients with primary nodal diffuse large B cell lymphoma. J Pathol. 2006;208:714–723. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

