Defining the antigen receptor-dependent regulatory network that induces arrest of cycling immature B-lymphocytes
- PMID: 21143896
- PMCID: PMC3004859
- DOI: 10.1186/1752-0509-4-169
Defining the antigen receptor-dependent regulatory network that induces arrest of cycling immature B-lymphocytes
Abstract
Background: Engagement of the antigen receptor on immature B-lymphocytes leads to cell cycle arrest, and subsequent apoptosis. This is an essential process for eliminating self reactive B cells during its different stages of development. However, the mechanism by which it is achieved is not completely understood.
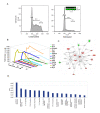
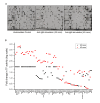
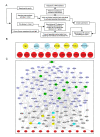
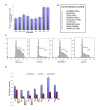
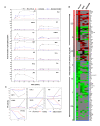
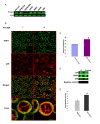
Results: Here we employed a systems biology approach that combined extensive experimentation with in silico methodologies to chart the network of receptor-activated pathways that mediated the arrest of immature B cells in the G1 phase of the cell cycle. Interestingly, we found that only a sparse network of signaling intermediates was recruited upon engagement of the antigen receptor. This then led to the activation of a restricted subset of transcription factors, with the consequent induction of genes primarily involved in the cell death pathway. Subsequent experiments revealed that the weak initiation of intracellular signaling pathways derived from desensitization of the receptor-proximal protein tyrosine kinase Lyn, to receptor-dependent activation. Intriguingly, the desensitization was a result of the constitutive activation of this kinase in unstimulated cells, which was likely maintained through a regulatory feedback loop involving the p38 MAP kinase. The high basal activity then attenuated the ability of the antigen receptor to recruit Lyn, and thereby also the downstream signaling intermediates. Finally, integration of these results into a mathematical model provided further substantiation to the novel finding that the ground state of the intracellular signaling machinery constitutes an important determinant of the outcome of receptor-induced cellular responses.
Conclusions: Our results identify the global events leading to the G1 arrest and subsequent apoptosis in immature B cells upon receptor activation.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

