A meiotic gene regulatory cascade driven by alternative fates for newly synthesized transcripts
- PMID: 21148298
- PMCID: PMC3016978
- DOI: 10.1091/mbc.E10-05-0448
A meiotic gene regulatory cascade driven by alternative fates for newly synthesized transcripts
Abstract
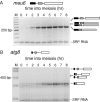
To determine the relative importance of transcriptional regulation versus RNA processing and turnover during the transition from proliferation to meiotic differentiation in the fission yeast Schizosaccharomyces pombe, we analyzed temporal profiles and effects of RNA surveillance factor mutants on expression of 32 meiotic genes. A comparison of nascent transcription with steady-state RNA accumulation reveals that the vast majority of these genes show a lag between maximal RNA synthesis and peak RNA accumulation. During meiosis, total RNA levels parallel 3' processing, which occurs in multiple, temporally distinct waves that peak from 3 to 6 h after meiotic induction. Most early genes and one middle gene, mei4, share a regulatory mechanism in which a specialized RNA surveillance factor targets newly synthesized transcripts for destruction. Mei4p, a member of the forkhead transcription factor family, in turn regulates a host of downstream genes. Remarkably, a spike in transcription is observed for less than one-third of the genes surveyed, and even these show evidence of RNA-level regulation. In aggregate, our findings lead us to propose that a regulatory cascade driven by changes in processing and stability of newly synthesized transcripts operates alongside the well-known transcriptional cascade as fission yeast cells enter meiosis.
Figures





References
-
- Àlvarez B, Moreno S. Fission yeast Tor2 promotes cell growth and represses cell differentiation. J Cell Sci. 2006;119:4475–4485. - PubMed
-
- Averbeck N, Sunder S, Sample N, Wise JA, Leatherwood J. Negative control contributes to an extensive program of meiotic splicing in fission yeast. Mol Cell. 2005;18:491–498. - PubMed
-
- Bähler J, Schuchert P, Grimm C, Kohli J. Synchronized meiosis and recombination in fission yeast: observations with pat1–114 diploid cells. Curr Genet. 1991;19:445–451. - PubMed
-
- Bühler M, Haas W, Gygi SP, Moazed D. RNA-dependent and -independent RNA turnover mechanisms contribute to heterochromatic gene silencing. Cell. 2007;129:707–721. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

