Overaccumulation of the chloroplast antisense RNA AS5 is correlated with decreased abundance of 5S rRNA in vivo and inefficient 5S rRNA maturation in vitro
- PMID: 21148395
- PMCID: PMC3022273
- DOI: 10.1261/rna.2336611
Overaccumulation of the chloroplast antisense RNA AS5 is correlated with decreased abundance of 5S rRNA in vivo and inefficient 5S rRNA maturation in vitro
Abstract
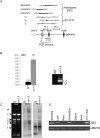
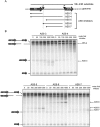
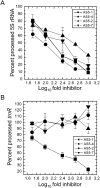
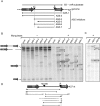
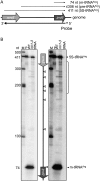
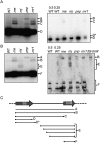
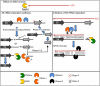
Post-transcriptional regulation in the chloroplast is exerted by nucleus-encoded ribonucleases and RNA-binding proteins. One of these ribonucleases is RNR1, a 3'-to-5' exoribonuclease of the RNase II family. We have previously shown that Arabidopsis rnr1-null mutants exhibit specific abnormalities in the expression of the rRNA operon, including the accumulation of precursor 23S, 16S, and 4.5S species and a concomitant decrease in the mature species. 5S rRNA transcripts, however, accumulate to a very low level in both precursor and mature forms, suggesting that they are unstable in the rnr1 background. Here we demonstrate that rnr1 plants overaccumulate an antisense RNA, AS5, that is complementary to the 5S rRNA, its intergenic spacer, and the downstream trnR gene, which encodes tRNA(Arg), raising the possibility that AS5 destabilizes 5S rRNA or its precursor and/or blocks rRNA maturation. To investigate this, we used an in vitro system that supports 5S rRNA and trnR processing. We show that AS5 inhibits 5S rRNA maturation from a 5S-trnR precursor, and shorter versions of AS5 demonstrate that inhibition requires intergenic sequences. To test whether the sense and antisense RNAs form double-stranded regions in vitro, treatment with the single-strand-specific mung bean nuclease was used. These results suggest that 5S-AS5 duplexes interfere with a sense-strand secondary structure near the endonucleolytic cleavage site downstream from the 5S rRNA coding region. We hypothesize that these duplexes are degraded by a dsRNA-specific ribonuclease in vivo, contributing to the 5S rRNA deficiency observed in rnr1.
Figures
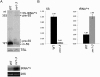


Similar articles
-
Overexpression of a natural chloroplast-encoded antisense RNA in tobacco destabilizes 5S rRNA and retards plant growth.BMC Plant Biol. 2010 Sep 29;10:213. doi: 10.1186/1471-2229-10-213. BMC Plant Biol. 2010. PMID: 20920268 Free PMC article.
-
RNR1, a 3'-5' exoribonuclease belonging to the RNR superfamily, catalyzes 3' maturation of chloroplast ribosomal RNAs in Arabidopsis thaliana.Nucleic Acids Res. 2005 May 12;33(8):2751-63. doi: 10.1093/nar/gki576. Print 2005. Nucleic Acids Res. 2005. PMID: 15891117 Free PMC article.
-
Maturation of atypical ribosomal RNA precursors in Helicobacter pylori.Nucleic Acids Res. 2019 Jun 20;47(11):5906-5921. doi: 10.1093/nar/gkz258. Nucleic Acids Res. 2019. PMID: 31006803 Free PMC article.
-
The cutting crew - ribonucleases are key players in the control of plastid gene expression.J Exp Bot. 2012 Feb;63(4):1663-73. doi: 10.1093/jxb/err401. Epub 2011 Dec 3. J Exp Bot. 2012. PMID: 22140236 Review.
-
Intermolecular base-paired interaction between complementary sequences present near the 3' ends of 5S rRNA and 18S (16S) rRNA might be involved in the reversible association of ribosomal subunits.Nucleic Acids Res. 1979 Dec 11;7(7):1913-29. doi: 10.1093/nar/7.7.1913. Nucleic Acids Res. 1979. PMID: 94160 Free PMC article. Review.
Cited by
-
Apoptosis-like programmed cell death induces antisense ribosomal RNA (rRNA) fragmentation and rRNA degradation in Leishmania.Cell Death Differ. 2012 Dec;19(12):1972-82. doi: 10.1038/cdd.2012.85. Epub 2012 Jul 6. Cell Death Differ. 2012. PMID: 22767185 Free PMC article.
-
Multifarious Evolutionary Pathways of a Nuclear RNA Editing Factor: Disjunctions in Coevolution of DOT4 and Its Chloroplast Target rpoC1eU488SL.Genome Biol Evol. 2019 Mar 1;11(3):798-813. doi: 10.1093/gbe/evz032. Genome Biol Evol. 2019. PMID: 30753430 Free PMC article.
-
RNase III, Ribosome Biogenesis and Beyond.Microorganisms. 2021 Dec 17;9(12):2608. doi: 10.3390/microorganisms9122608. Microorganisms. 2021. PMID: 34946208 Free PMC article. Review.
-
DNA:RNA Hybrids Are Major Dinoflagellate Minicircle Molecular Types.Int J Mol Sci. 2023 Jun 2;24(11):9651. doi: 10.3390/ijms24119651. Int J Mol Sci. 2023. PMID: 37298602 Free PMC article.
-
Oryza sativa ObgC1 Acts as a Key Regulator of DNA Replication and Ribosome Biogenesis in Chloroplast Nucleoids.Rice (N Y). 2021 Jul 12;14(1):65. doi: 10.1186/s12284-021-00498-5. Rice (N Y). 2021. PMID: 34251486 Free PMC article.
References
-
- Andrade JM, Pobre Vn, Silva IJ, Domingues S, Arraiano CM, Ciaran C. 2009. The role of 3′–5′ exoribonucleases in RNA degradation. In Progress in molecular biology and translational science, Vol. 85, pp. 187–229. Academic Press, New York. - PubMed
-
- Bollenbach TJ, Sharwood RE, Gutierrez R, Lerbs-Mache S, Stern DB 2009. The RNA-binding proteins CSP41a and CSP41b may regulate transcription and translation of chloroplast-encoded RNAs in Arabidopsis. Plant Mol Biol 69: 541–552 - PubMed
-
- Britton RA, Wen T, Schaefer L, Pellegrini O, Uicker WC, Mathy N, Tobin C, Daou R, Szyk J, Condon C 2007. Maturation of the 5′ end of Bacillus subtilis 16S rRNA by the essential ribonuclease YkqC/RNase J1. Mol Microbiol 63: 127–138 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
