A thymus-specific noncoding RNA, Thy-ncR1, is a cytoplasmic riboregulator of MFAP4 mRNA in immature T-cell lines
- PMID: 21162727
- PMCID: PMC3023731
- DOI: 10.1186/1471-2199-11-99
A thymus-specific noncoding RNA, Thy-ncR1, is a cytoplasmic riboregulator of MFAP4 mRNA in immature T-cell lines
Abstract
Background: Postgenomic transcriptome analyses have identified large numbers of noncoding (nc)RNAs in mammalian cells. However, the biological function of long ncRNAs in mammalian cells remains largely unknown. Our recent expression profiling of selected human long ncRNAs revealed that a majority were expressed in an organ-specific manner, suggesting their function was linked to specific physiological phenomena in each organ. We investigated the characteristics and function of ncRNAs that were specifically expressed in the thymus, the site of T-cell selection and maturation.
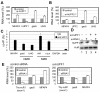
Results: Expression profiling of 10 thymus-specific ncRNAs in 17 T-cell leukemia cell lines derived from various stages of T-cell maturation revealed that HIT14168 ncRNA, named Thy-ncR1, was specifically expressed in cell lines derived from stage III immature T cells in which the neighbouring CD1 gene cluster is also specifically activated. The Thy-ncR1 precursor exhibited complex alternative splicing patterns and differential usage of the 5' terminus leading to the production of an estimated 24 isoforms, which were predominantly located in the cytoplasm. Selective RNAi knockdown of each Thy-ncR1 isoform demonstrated that microfibril-associated glycoprotein 4 (MFAP4) mRNA was negatively regulated by two major Thy-ncR1 isoforms. Intriguingly, the MFAP4 mRNA level was controlled by a hUPF1-dependent mRNA degradation pathway in the cytoplasm distinct from nonsense-mediated decay.
Conclusions: This study identified Thy-ncR1 ncRNA to be specifically expressed in stage III immature T cells in which the neighbouring CD1 gene cluster was activated. Complex alternative splicing produces multiple Thy-ncR1 isoforms. Two major Thy-ncR1 isoforms are cytoplasmic riboregulators that suppress the expression of MFAP4 mRNA, which is degraded by an uncharacterized hUPF1-dependent pathway.
Figures





References
-
- Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, Kodzius R, Shimokawa K, Bajic VB, Brenner SE, Batalov S, Forrest AR, Zavolan M, Davis MJ, Wilming LG, Aidinis V, Allen JE, Ambesi-Impiombato A, Apweiler R, Aturaliya RN, Bailey TL, Bansal M, Baxter L, Beisel KW, Bersano T, Bono H. FANTOM Consortium, RIKEN Genome Exploration Research Group and Genome Science Group (Genome Network Project Core Group) et al. The transcriptional landscape of the mammalian genome. Science. 2005;309:1559–63. doi: 10.1126/science.1112014. - DOI - PubMed
-
- Cheng J, Kapranov P, Drenkow J, Dike S, Brubaker S, Patel S, Long J, Stern D, Tammana H, Helt G, Sementchenko V, Piccolboni A, Bekiranov S, Bailey DK, Ganesh M, Ghosh S, Bell I, Gerhard DS, Gingeras TR. Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science. 2005;308:1149–54. doi: 10.1126/science.1108625. - DOI - PubMed
-
- ENCODE Project Consortium, Birney E, Stamatoyannopoulos JA, Dutta A, Guigó R, Gingeras TR, Margulies EH, Weng Z, Snyder M, Dermitzakis ET, Thurman RE, Kuehn MS, Taylor CM, Neph S, Koch CM, Asthana S, Malhotra A, Adzhubei I, Greenbaum JA, Andrews RM, Flicek P, Boyle PJ, Cao H, Carter NP, Clelland GK, Davis S, Day N, Dhami P, Dillon SC, Dorschner MO, Fiegler H. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature. 2007;447:799–816. doi: 10.1038/nature05874. - DOI - PMC - PubMed
-
- Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermüller J, Hofacker IL, Bell I, Cheung E, Drenkow J, Dumais E, Patel S, Helt G, Ganesh M, Ghosh S, Piccolboni A, Sementchenko V, Tammana H, Gingeras TR. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science. 2007;316:1484–8. doi: 10.1126/science.1138341. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

