Multiplex target enrichment using DNA indexing for ultra-high throughput SNP detection
- PMID: 21163834
- PMCID: PMC3041504
- DOI: 10.1093/dnares/dsq029
Multiplex target enrichment using DNA indexing for ultra-high throughput SNP detection
Abstract
Screening large numbers of target regions in multiple DNA samples for sequence variation is an important application of next-generation sequencing but an efficient method to enrich the samples in parallel has yet to be reported. We describe an advanced method that combines DNA samples using indexes or barcodes prior to target enrichment to facilitate this type of experiment. Sequencing libraries for multiple individual DNA samples, each incorporating a unique 6-bp index, are combined in equal quantities, enriched using a single in-solution target enrichment assay and sequenced in a single reaction. Sequence reads are parsed based on the index, allowing sequence analysis of individual samples. We show that the use of indexed samples does not impact on the efficiency of the enrichment reaction. For three- and nine-indexed HapMap DNA samples, the method was found to be highly accurate for SNP identification. Even with sequence coverage as low as 8x, 99% of sequence SNP calls were concordant with known genotypes. Within a single experiment, this method can sequence the exonic regions of hundreds of genes in tens of samples for sequence and structural variation using as little as 1 μg of input DNA per sample.
Figures


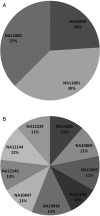
References
-
- Dahl F., Gullberg M., Stenberg J., Landegren U., Nilsson M. Multiplex amplification enabled by selective circularization of large sets of genomic DNA fragments. Nucleic Acids Res. 2005;33:e71. doi:10.1093/nar/gni070. - DOI - PMC - PubMed
-
- Landegren U., Schallmeiner E., Nilsson M., et al. Molecular tools for a molecular medicine: analyzing genes, transcripts and proteins using padlock and proximity probes. J. Mol. Recognit. 2004;17:194–7. doi:10.1002/jmr.664. - DOI - PubMed
-
- Li J.B., Gao Y., Aach J., et al. Multiplex padlock targeted sequencing reveals human hypermutable CpG variations. Genome Res. 2009;19:1606–15. doi:10.1101/gr.092213.109. - DOI - PMC - PubMed
-
- Lovett M., Kere J., Hinton L.M. Direct selection: a method for the isolation of cDNAs encoded by large genomic regions. Proc. Natl Acad. Sci. USA. 1991;88:9628–32. doi:10.1073/pnas.88.21.9628. - DOI - PMC - PubMed
-
- Porreca G.J., Zhang K., Li J.B., et al. Multiplex amplification of large sets of human exons. Nat. Methods. 2007;4:931–6. doi:10.1038/nmeth1110. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

