Mammalian MTHFD2L encodes a mitochondrial methylenetetrahydrofolate dehydrogenase isozyme expressed in adult tissues
- PMID: 21163947
- PMCID: PMC3037629
- DOI: 10.1074/jbc.M110.196840
Mammalian MTHFD2L encodes a mitochondrial methylenetetrahydrofolate dehydrogenase isozyme expressed in adult tissues
Abstract
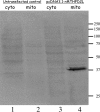
Previous studies in our laboratory showed that isolated, intact adult rat liver mitochondria are able to oxidize the 3-carbon of serine and the N-methyl carbon of sarcosine to formate without the addition of any other cofactors or substrates. Conversion of these 1-carbon units to formate requires several folate-interconverting enzymes in mitochondria. The enzyme(s) responsible for conversion of 5,10-methylene-tetrahydrofolate (CH(2)-THF) to 10-formyl-THF in adult mammalian mitochondria are currently unknown. A new mitochondrial CH(2)-THF dehydrogenase isozyme, encoded by the MTHFD2L gene, has now been identified. The recombinant protein exhibits robust NADP(+)-dependent CH(2)-THF dehydrogenase activity when expressed in yeast. The enzyme is localized to mitochondria when expressed in CHO cells and behaves as a peripheral membrane protein, tightly associated with the matrix side of the mitochondrial inner membrane. The MTHFD2L gene is subject to alternative splicing and is expressed in adult tissues in humans and rodents. This CH(2)-THF dehydrogenase isozyme thus fills the remaining gap in the pathway from CH(2)-THF to formate in adult mammalian mitochondria.
Figures

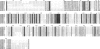


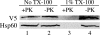



References
-
- Appling D. R. (1991) FASEB J. 5, 2645–2651 - PubMed
-
- Woeller C. F., Anderson D. D., Szebenyi D. M., Stover P. J. (2007) J. Biol. Chem. 282, 17623–17631 - PubMed
-
- Davis S. R., Stacpoole P. W., Williamson J., Kick L. S., Quinlivan E. P., Coats B. S., Shane B., Bailey L. B., Gregory J. F., 3rd. (2004) Am. J. Physiol. Endocrinol. Metab. 286, E272–EE279 - PubMed
-
- Fu T. F., Rife J. P., Schirch V. (2001) Arch. Biochem. Biophys. 393, 42–50 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

