TDRD3 is an effector molecule for arginine-methylated histone marks
- PMID: 21172665
- PMCID: PMC3090733
- DOI: 10.1016/j.molcel.2010.11.024
TDRD3 is an effector molecule for arginine-methylated histone marks
Abstract
Specific sites of histone tail methylation are associated with transcriptional activity at gene loci. These methyl marks are interpreted by effector molecules, which harbor protein domains that bind the methylated motifs and facilitate either active or inactive states of transcription. CARM1 and PRMT1 are transcriptional coactivators that deposit H3R17me2a and H4R3me2a marks, respectively. We used a protein domain microarray approach to identify the Tudor domain-containing protein TDRD3 as a "reader" of these marks. Importantly, TDRD3 itself is a transcriptional coactivator. This coactivator activity requires an intact Tudor domain. TDRD3 is recruited to an estrogen-responsive element in a CARM1-dependent manner. Furthermore, ChIP-seq analysis of TDRD3 reveals that it is predominantly localized to transcriptional start sites. Thus, TDRD3 is an effector molecule that promotes transcription by binding methylarginine marks on histone tails.
Copyright © 2010 Elsevier Inc. All rights reserved.
Figures


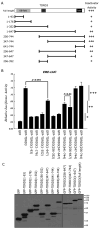

References
-
- An W, Kim J, Roeder RG. Ordered cooperative functions of PRMT1, p300, and CARM1 in transcriptional activation by p53. Cell. 2004;117:735–748. - PubMed
-
- Bannister AJ, Zegerman P, Partridge JF, Miska EA, Thomas JO, Allshire RC, Kouzarides T. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature. 2001;410:120–124. - PubMed
-
- Chen D, Ma H, Hong H, Koh SS, Huang SM, Schurter BT, Aswad DW, Stallcup MR. Regulation of transcription by a protein methyltransferase. Science. 1999;284:2174–2177. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

