Widespread regulatory activity of vertebrate microRNA* species
- PMID: 21177881
- PMCID: PMC3022280
- DOI: 10.1261/rna.2537911
Widespread regulatory activity of vertebrate microRNA* species
Abstract
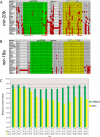
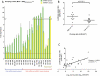
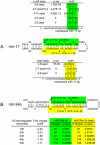
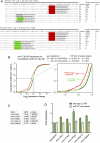
An obligate intermediate during microRNA (miRNA) biogenesis is an ~22-nucleotide RNA duplex, from which the mature miRNA is preferentially incorporated into a silencing complex. Its partner miRNA* species is generally regarded as a passenger RNA, whose regulatory capacity has not been systematically examined in vertebrates. Our bioinformatic analyses demonstrate that a substantial fraction of miRNA* species are stringently conserved over vertebrate evolution, collectively exhibit greatest conservation in their seed regions, and define complementary motifs whose conservation across vertebrate 3'-UTR evolution is statistically significant. Functional tests of 22 miRNA expression constructs revealed that a majority could repress both miRNA and miRNA* perfect match reporters, and the ratio of miRNA:miRNA* sensor repression was correlated with the endogenous ratio of miRNA:miRNA* reads. Analysis of microarray data provided transcriptome-wide evidence for the regulation of seed-matched targets for both mature and star strand species of several miRNAs relevant to oncogenesis, including mir-17, mir-34a, and mir-19. Finally, 3'-UTR sensor assays and mutagenesis tests confirmed direct repression of five miR-19* targets via star seed sites. Overall, our data demonstrate that miRNA* species have demonstrable impact on vertebrate regulatory networks and should be taken into account in studies of miRNA functions and their contribution to disease states.
Figures

Similar articles
-
The fate of miRNA* strand through evolutionary analysis: implication for degradation as merely carrier strand or potential regulatory molecule?PLoS One. 2010 Jun 30;5(6):e11387. doi: 10.1371/journal.pone.0011387. PLoS One. 2010. PMID: 20613982 Free PMC article.
-
Functional conservation of both CDS- and 3'-UTR-located microRNA binding sites between species.Mol Biol Evol. 2015 Mar;32(3):623-8. doi: 10.1093/molbev/msu323. Epub 2014 Nov 19. Mol Biol Evol. 2015. PMID: 25414126
-
New class of microRNA targets containing simultaneous 5'-UTR and 3'-UTR interaction sites.Genome Res. 2009 Jul;19(7):1175-83. doi: 10.1101/gr.089367.108. Epub 2009 Mar 31. Genome Res. 2009. PMID: 19336450 Free PMC article.
-
miRNA Targeting: Growing beyond the Seed.Trends Genet. 2019 Mar;35(3):215-222. doi: 10.1016/j.tig.2018.12.005. Epub 2019 Jan 9. Trends Genet. 2019. PMID: 30638669 Free PMC article. Review.
-
Functional relevance of miRNA sequences in human disease.Mutat Res. 2012 Mar 1;731(1-2):14-9. doi: 10.1016/j.mrfmmm.2011.10.014. Epub 2011 Nov 6. Mutat Res. 2012. PMID: 22085809 Free PMC article. Review.
Cited by
-
Computational Prediction of miRNA Genes from Small RNA Sequencing Data.Front Bioeng Biotechnol. 2015 Jan 26;3:7. doi: 10.3389/fbioe.2015.00007. eCollection 2015. Front Bioeng Biotechnol. 2015. PMID: 25674563 Free PMC article. Review.
-
The impact of age, biogenesis, and genomic clustering on Drosophila microRNA evolution.RNA. 2013 Sep;19(9):1295-308. doi: 10.1261/rna.039248.113. Epub 2013 Jul 23. RNA. 2013. PMID: 23882112 Free PMC article.
-
Comprehensive re-analysis of hairpin small RNAs in fungi reveals loci with conserved links.Elife. 2022 Dec 9;11:e83691. doi: 10.7554/eLife.83691. Elife. 2022. PMID: 36484778 Free PMC article.
-
One locus, several functional RNAs-emerging roles of the mechanisms responsible for the sequence variability of microRNAs.Biol Futur. 2023 Jun;74(1-2):17-28. doi: 10.1007/s42977-023-00154-7. Epub 2023 Feb 27. Biol Futur. 2023. PMID: 36847925 Review.
-
The role of miRNA and lncRNA in heterotopic ossification pathogenesis.Stem Cell Res Ther. 2022 Dec 15;13(1):523. doi: 10.1186/s13287-022-03213-3. Stem Cell Res Ther. 2022. PMID: 36522666 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
