Meganucleases and other tools for targeted genome engineering: perspectives and challenges for gene therapy
- PMID: 21182466
- PMCID: PMC3267165
- DOI: 10.2174/156652311794520111
Meganucleases and other tools for targeted genome engineering: perspectives and challenges for gene therapy
Abstract
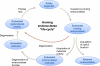
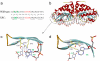
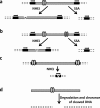
The importance of safer approaches for gene therapy has been underscored by a series of severe adverse events (SAEs) observed in patients involved in clinical trials for Severe Combined Immune Deficiency Disease (SCID) and Chromic Granulomatous Disease (CGD). While a new generation of viral vectors is in the process of replacing the classical gamma-retrovirus-based approach, a number of strategies have emerged based on non-viral vectorization and/or targeted insertion aimed at achieving safer gene transfer. Currently, these methods display lower efficacies than viral transduction although many of them can yield more than 1% of engineered cells in vitro. Nuclease-based approaches, wherein an endonuclease is used to trigger site-specific genome editing, can significantly increase the percentage of targeted cells. These methods therefore provide a real alternative to classical gene transfer as well as gene editing. However, the first endonuclease to be in clinic today is not used for gene transfer, but to inactivate a gene (CCR5) required for HIV infection. Here, we review these alternative approaches, with a special emphasis on meganucleases, a family of naturally occurring rare-cutting endonucleases, and speculate on their current and future potential.
Figures

References
-
- Cavazzana-Calvo M, Hacein-Bey S, de Saint Basile G, et al. Gene therapy of human severe combined immunodeficiency (SCID)-X1 disease. Science. 2000;288:669–72. - PubMed
-
- Gaspar HB, Parsley KL, Howe S, et al. Gene therapy of X-linked severe combined immunodeficiency by use of a pseudotyped gammaretroviral vector. Lancet. 2004;364:2181–7. - PubMed
-
- Hacein-Bey-Abina S, Von Kalle C, Schmidt M, et al. LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1. Science. 2003;302:415–9. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

