Activity map of the tammar X chromosome shows that marsupial X inactivation is incomplete and escape is stochastic
- PMID: 21182760
- PMCID: PMC3046482
- DOI: 10.1186/gb-2010-11-12-r122
Activity map of the tammar X chromosome shows that marsupial X inactivation is incomplete and escape is stochastic
Abstract
Background: X chromosome inactivation is a spectacular example of epigenetic silencing. In order to deduce how this complex system evolved, we examined X inactivation in a model marsupial, the tammar wallaby (Macropus eugenii). In marsupials, X inactivation is known to be paternal, incomplete and tissue-specific, and occurs in the absence of an XIST orthologue.
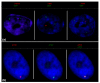
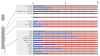
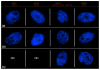
Results: We examined expression of X-borne genes using quantitative PCR, revealing a range of dosage compensation for different loci. To assess the frequency of 1X- or 2X-active fibroblasts, we investigated expression of 32 X-borne genes at the cellular level using RNA-FISH. In female fibroblasts, two-color RNA-FISH showed that genes were coordinately expressed from the same X (active X) in nuclei in which both loci were inactivated. However, loci on the other X escape inactivation independently, with each locus showing a characteristic frequency of 1X-active and 2X-active nuclei, equivalent to stochastic escape. We constructed an activity map of the tammar wallaby inactive X chromosome, which identified no relationship between gene location and extent of inactivation, nor any correlation with the presence or absence of a Y-borne paralog.
Conclusions: In the tammar wallaby, one X (presumed to be maternal) is expressed in all cells, but genes on the other (paternal) X escape inactivation independently and at characteristic frequencies. The paternal and incomplete X chromosome inactivation in marsupials, with stochastic escape, appears to be quite distinct from the X chromosome inactivation process in eutherians. We find no evidence for a polar spread of inactivation from an X inactivation center.
Figures






References
-
- Page J, Berrios S, Rufas JS, Parra MT, Sija JA, Heyting C, Fernandez-Donosoo R. The meiotic pairing of X and Y chromosomes in the marsupial species Thylamys elegans is maintained by a dense plate developed from their axial elements. J Cell Sci. 2003;116:551–560. doi: 10.1242/jcs.00252. - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

