Hybridization and restricted gene flow between native and introduced stocks of Alpine whitefish (Coregonus sp.) across multiple environments
- PMID: 21199024
- PMCID: PMC3045663
- DOI: 10.1111/j.1365-294X.2010.04961.x
Hybridization and restricted gene flow between native and introduced stocks of Alpine whitefish (Coregonus sp.) across multiple environments
Abstract
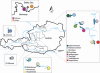
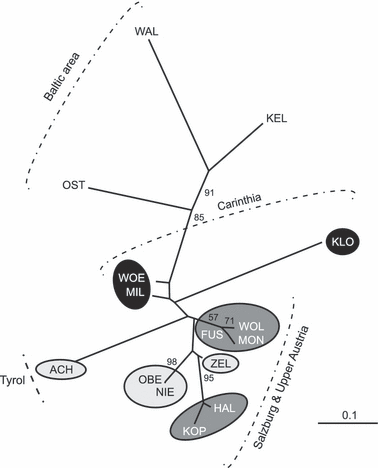
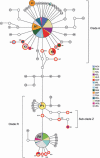
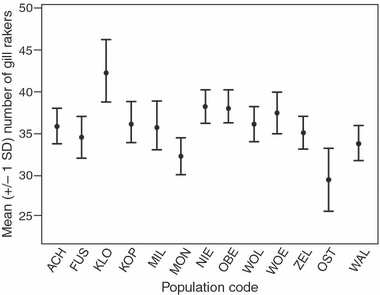
Translocations of Baltic whitefish (Coregonus sp.) into Austrian Alpine lakes have created 'artificial hybrid zones', threatening the genetic integrity of native lineages. We evaluate the genetic structure of Coregonus in Austrian lakes and characterize hybridization and introgression between native and introduced lineages. Fifteen populations (N=747) were assessed for allelic variation at eight microsatellite loci and a reduced set (N=253) for variation across two mtDNA genes (cyt b and NADH-3). Bayesian approaches were used to estimate individual admixture proportions (q-values) and classify genotypes as native, introduced or hybrids. q-value distributions varied among populations highlighting differential hybridization and introgression histories. Many lakes revealed a clear distinction between native and introduced genotypes despite hybridization, whereas some locations revealed hybrid swarms. Genetic structure among lakes was congruent with morphological divergence and novelty raising speculation of multiple taxa, including a population south of the Alps, outside the putative native range of Coregonus. Although statistically congruent with inferences based on nuclear markers, mitochondrial haplotype data was not diagnostic with respect to native and non-native lineages, supporting that the Alpine region was colonized post-glacially by an admixture of mtDNA lineages, which coalesce >1 Ma. Mechanisms promoting or eroding lineage isolation are discussed, as well as a high potential to conserve native Alpine lineages despite the extensive historical use of introduced Baltic stocks.
© 2010 Blackwell Publishing Ltd.
Figures

Similar articles
-
Contrasting patterns of mitochondrial DNA and microsatellite introgressive hybridization between lineages of lake whitefish (Coregonus clupeaformis); relevance for speciation.Mol Ecol. 2001 Apr;10(4):965-85. doi: 10.1046/j.1365-294x.2001.01252.x. Mol Ecol. 2001. PMID: 11348504
-
Species introduction promotes hybridization and introgression in Coregonus: is there sign of selection against hybrids?Mol Ecol. 2011 Sep;20(18):3838-55. doi: 10.1111/j.1365-294X.2011.05209.x. Epub 2011 Aug 10. Mol Ecol. 2011. PMID: 21831252
-
Testing the devil's impact on southern Baltic and North Sea basins whitefish (Coregonus spp.) diversity.BMC Evol Biol. 2018 Dec 29;18(1):208. doi: 10.1186/s12862-018-1339-2. BMC Evol Biol. 2018. PMID: 30594141 Free PMC article.
-
Genetic population structure of sympatric and allopatric populations of Baltic ciscoes (Coregonus albula complex, Teleostei, Coregonidae).BMC Evol Biol. 2010 Mar 29;10:85. doi: 10.1186/1471-2148-10-85. BMC Evol Biol. 2010. PMID: 20350300 Free PMC article.
-
Hybrids and horizontal transfer: introgression allows adaptive allele discovery.J Exp Bot. 2017 Nov 28;68(20):5453-5470. doi: 10.1093/jxb/erx297. J Exp Bot. 2017. PMID: 29096001 Review.
Cited by
-
Understanding admixture patterns in supplemented populations: a case study combining molecular analyses and temporally explicit simulations in Atlantic salmon.Evol Appl. 2013 Feb;6(2):218-30. doi: 10.1111/j.1752-4571.2012.00280.x. Epub 2012 Jun 14. Evol Appl. 2013. PMID: 23798972 Free PMC article.
-
First insight into the phylogeny of fine-leaved Festuca in the Altai Mountain Country based on genome-wide genotyping.Ecol Evol. 2023 Apr 2;13(4):e9943. doi: 10.1002/ece3.9943. eCollection 2023 Apr. Ecol Evol. 2023. PMID: 37021080 Free PMC article.
-
Genetic variation in brown trout Salmo trutta across the Danube, Rhine, and Elbe headwaters: a failure of the phylogeographic paradigm?BMC Evol Biol. 2013 Aug 26;13:176. doi: 10.1186/1471-2148-13-176. BMC Evol Biol. 2013. PMID: 23972037 Free PMC article.
-
Within-range translocations and their consequences in European larch.PLoS One. 2015 May 22;10(5):e0127516. doi: 10.1371/journal.pone.0127516. eCollection 2015. PLoS One. 2015. PMID: 26000791 Free PMC article.
-
Anthropogenic hybridization between endangered migratory and commercially harvested stationary whitefish taxa (Coregonus spp.).Evol Appl. 2014 Nov;7(9):1068-83. doi: 10.1111/eva.12166. Epub 2014 Jul 8. Evol Appl. 2014. PMID: 25553068 Free PMC article.
References
-
- Allendorf FW, Leary RF, Spruell P, Wenburg JK. The problems with hybrids: setting conservation guidelines. Trends in Ecology and Evolution. 2001;16:613–622.
-
- Bandelt HJ, Forster P, Röhl A. Median-joining networks for inferring intraspecific phylogenies. Molecular Biology and Evolution. 1999;16:37–48. http://www.fluxus-engineering.com. - PubMed
-
- Belkhir K, Borsa P, Chikhi L, Raufaste N, Bonhomme F. GENETIX 4.05, logiciel sous Windows TM pour la génétique des populations′. Montpellier, France: Laboratoire Génome, Populations, Interactions, CNRS UMR 5171, Université de Montpellier II; 1996. 2004.
-
- Benda H. Der Riedling des Traunsees. Österreichs Fischerei. 1949;2:219–221.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

