Structural basis for scaffolding-mediated assembly and maturation of a dsDNA virus
- PMID: 21220301
- PMCID: PMC3029737
- DOI: 10.1073/pnas.1015739108
Structural basis for scaffolding-mediated assembly and maturation of a dsDNA virus
Abstract
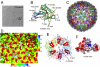
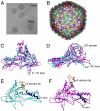
Formation of many dsDNA viruses begins with the assembly of a procapsid, containing scaffolding proteins and a multisubunit portal but lacking DNA, which matures into an infectious virion. This process, conserved among dsDNA viruses such as herpes viruses and bacteriophages, is key to forming infectious virions. Bacteriophage P22 has served as a model system for this study in the past several decades. However, how capsid assembly is initiated, where and how scaffolding proteins bind to coat proteins in the procapsid, and the conformational changes upon capsid maturation still remain elusive. Here, we report Cα backbone models for the P22 procapsid and infectious virion derived from electron cryomicroscopy density maps determined at 3.8- and 4.0-Å resolution, respectively, and the first procapsid structure at subnanometer resolution without imposing symmetry. The procapsid structures show the scaffolding protein interacting electrostatically with the N terminus (N arm) of the coat protein through its C-terminal helix-loop-helix motif, as well as unexpected interactions between 10 scaffolding proteins and the 12-fold portal located at a unique vertex. These suggest a critical role for the scaffolding proteins both in initiating the capsid assembly at the portal vertex and propagating its growth on a T = 7 icosahedral lattice. Comparison of the procapsid and the virion backbone models reveals coordinated and complex conformational changes. These structural observations allow us to propose a more detailed molecular mechanism for the scaffolding-mediated capsid assembly initiation including portal incorporation, release of scaffolding proteins upon DNA packaging, and maturation into infectious virions.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Earnshaw WC, Hendrix RW, King J. Structural studies of bacteriophage lambda heads and proheads by small angle X-ray diffraction. J Mol Biol. 1979;134:575–594. - PubMed
-
- King J, Chiu W. The procapsid-to-capsid transition in double-stranded DNA bacteriophages. In: Chiu W, Burnett RM, Garcea RL, editors. Structural Biology of Viruses. New York: Oxford University Press; 1997. pp. 288–311.
-
- Heymann JB, et al. Dynamics of herpes simplex virus capsid maturation visualized by time-lapse cryo-electron microscopy. Nat Struct Biol. 2003;10:334–341. - PubMed
-
- Bamford DH, Grimes JM, Stuart DI. What does structure tell us about virus evolution? Curr Opin Struct Biol. 2005;15:655–663. - PubMed
-
- Caspar DL, Klug A. Physical principles in the construction of regular viruses. Cold Spring Harbor Symp Quant Biol. 1962;27:1–24. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

