Substrate specificity and catalysis by the editing active site of Alanyl-tRNA synthetase from Escherichia coli
- PMID: 21241052
- PMCID: PMC3069921
- DOI: 10.1021/bi1013535
Substrate specificity and catalysis by the editing active site of Alanyl-tRNA synthetase from Escherichia coli
Abstract
Aminoacyl-tRNA synthetases (ARSs) enhance the fidelity of protein synthesis through multiple mechanisms, including hydrolysis of the adenylate and cleavage of misacylated tRNA. Alanyl-tRNA synthetase (AlaRS) limits misacylation with glycine and serine by use of a dedicated editing domain, and a mutation in this activity has been genetically linked to a mouse model of a progressive neurodegenerative disease. Using the free-standing Pyrococcus horikoshii AlaX editing domain complexed with serine as a model and both Ser-tRNA(Ala) and Ala-tRNA(Ala) as substrates, the deacylation activities of the wild type and five different Escherichia coli AlaRS editing site substitution mutants were characterized. The wild-type AlaRS editing domain deacylated Ser-tRNA(Ala) with a k(cat)/K(M) of 6.6 × 10(5) M(-1) s(-1), equivalent to a rate enhancement of 6000 over the rate of enzyme-independent deacylation but only 12.2-fold greater than the rate with Ala-tRNA(Ala). While the E664A and T567G substitutions only minimally decreased k(cat)/K(M,) Q584H, I667E, and C666A AlaRS were more compromised in activity, with decreases in k(cat)/K(M) in the range of 6-, 6.6-, and 15-fold. C666A AlaRS was 1.7-fold more active on Ala-tRNA(Ala) relative to Ser-tRNA(Ala), providing the only example of a true reversal of substrate specificity and highlighting a potential role of the coordinated zinc in editing substrate specificity. Along with the potentially serious physiological consequences of serine misincorporation, the relatively modest specificity of the AlaRS editing domain may provide a rationale for the widespread phylogenetic distribution of AlaX free-standing editing domains, thereby contributing a further mechanism to lower concentrations of misacylated tRNA(Ala).
Figures




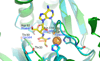
References
-
- Ibba M, Francklyn C, Cusack S, editors. The Aminoacyl-tRNA Synthetases. Landes Bioscience: Georgetown, Texas; 2005.
-
- Murakami H, Ohta A, Suga H. Bases in the anticodon loop of tRNA(Ala)(GGC) prevent misreading. Nat. Struct. Mol. Biol. 2009;16:353–358. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

