Dynamic histone acetylation is critical for cotranscriptional spliceosome assembly and spliceosomal rearrangements
- PMID: 21245291
- PMCID: PMC3033250
- DOI: 10.1073/pnas.1011982108
Dynamic histone acetylation is critical for cotranscriptional spliceosome assembly and spliceosomal rearrangements
Abstract
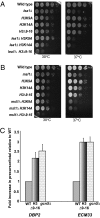
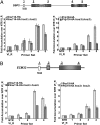
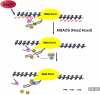
Assembly of the spliceosome onto pre-mRNA is a dynamic process involving the ordered exchange of snRNPs to form the catalytically active spliceosome. These ordered rearrangements have recently been shown to occur cotranscriptionally, while the RNA polymerase is still actively engaged with the chromatin template. We previously demonstrated that the histone acetyltransferase Gcn5 is required for U2 snRNP association with the branchpoint. Here we provide evidence that histone acetylation and deacetylation facilitate proper cotranscriptional association of spliceosomal snRNPs. As with GCN5, mutation or deletion of Gcn5-targeted histone H3 residues leads to synthetic lethality when combined with deletion of the genes encoding the U2 snRNP components Lea1 or Msl1. Gcn5 associates throughout intron-containing genes and, in the absence of the histone deacetylases Hos3 and Hos2, enhanced histone H3 acetylation is observed throughout the body of genes. Deletion of histone deacetylaces also results in persistent association of the U2 snRNP and a severe defect in the association of downstream factors. These studies show that cotranscriptional spliceosome rearrangements are driven by dynamic changes in the acetylation state of histones and provide a model whereby yeast spliceosome assembly is tightly coupled to histone modification.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

